
- 352 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Alkaliphiles
About this book
One of the most fascinating aspects of alkaliphiles is their ability to maintain pH homeostasis under extreme environmental conditions. This work provides a treatment of alkaliphilic microbiology, supported by molecular studies on the genetics of alkaliphilic "Bacillus" strain. Genomic analysis of "Bacillus halodurans" C-125 has been started and the genes responsible for alkaliphily are described. In addition to a basic background of alkaliphiles, including discussions of cell structures, physiology and molecular biology, "Alkaliphiles" presents an analysis of extracellular enzymes. Research on numerous enzymes including alkaline proteases, starch-degrading enzymes, cellulases, mannan-degrading enzymes, and many others is described in depth with relevant industrial applications.
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
7 Extracellular Enzymes
7.1 Isolation Procedures for Alkaline Enzymes
Enzymes having pH optimum for enzyme action in the alkaline regions are called alkaline enzymes. These include, for example, alkaline proteases (pH 10-12), alkaline amylases (pH 9-11), alkaline cellulases (pH 8-10), and others. Except for alkaline proteases these enzymes had not been isolated before our rediscovery of alkaliphilic microorganisms in 1968. Isolation of alkaline enzymes means the isolation of alkaliphilic microorganisms producing alkaline enzymes. Many screening procedures for these microorganisms have been developed. General procedures are described below with specific methods given in subsequent sections.
7.1.1 Protease Activity
Polypeptone in Horikoshi-I or -II medium is substituted for crude casein such as skim milk (1% w/v) and microorganisms are inoculated on the solidified media. After two or three days incubation at 37 °C, peptonization activity is directly observed as colonies hydrolyzing casein surrounded by a white halo. Candidates producing protease are cultured in the liquid media described in Table 7.1, and activity in the culture media are determined by conventional methods. Semi-quantitative assay using pulp disks is also a good method.
| Ingredients (g/l) | Protease | Amylase | Cellulase | β-1,3-Glucanase | ||||
|---|---|---|---|---|---|---|---|---|
| Soluble starch | 50 | 40 | 15 | 80 | 80 | |||
| CMC | 20 | |||||||
| Pachyman | 20 | |||||||
| Soya meal | 20 | 30 | 30 | 20 | ||||
| Ground barley | 50 | 100 | 100 | |||||
| Soya bean extract | 10 | 10 | ||||||
| Polypeptone | 10 | 20 | 5 | 5 | ||||
| Fish extract | 20 | |||||||
| Na-caseinate | 10 | |||||||
| Soya bean oil | 5.5 | |||||||
| Polyglycol | 0.1 | 0.1 | ||||||
| Yeast extract | 5 | 5 | ||||||
| KH2PO4 | 1 | |||||||
| Na2HPO4•10H2O | 9 | 9 | ||||||
| NaCl | 2.5 | |||||||
| CaCl2 | 1 | |||||||
| CaCO3 | 5 | |||||||
| MgSO4 | 0.2 | 0.2 | ||||||
| Na2CO3 | 10 | 10 | 10 | 10 | 10 | 10 | ||
| NaHCO3 | 10 | |||||||
7.1.2 Amylase Activity
Isolated microorganisms are grown in solidified Horikoshi-II medium. After two to three days incubation at 37 °C, iodine solution is poured onto plates. The colonies surrounded by a white halo are the amylase producers.
7.1.3 Cellulase Activity
Instead of glucose, 1% CMC is added to the Horikoshi-I medium solidified by agar containing 0.03% w/v Congo red, and soil samples spread on the plates. After two to three days incubation at 35-40 °C, white halos form around bacterial colonies producing CMCase.
7.1.4 β-1,3-Glucanase
Defatted pachyman powder is used as the β-1,3-glucan. Laminaran is also a good substrate for isolating microorganisms. Congo red (about 0.01-0.03% w/v) is added to the
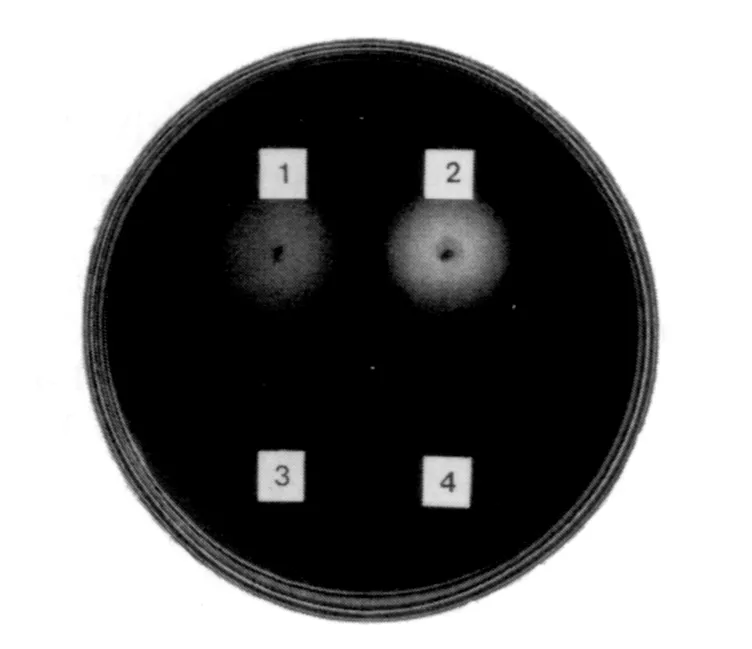
media described above to indicate β-1,3-glucanase activity. The colonies surrounded by white halos are the enzyme producers. Pachyman should be defatted by ether, otherwise very poor growth of microorganisms is observed due to the presence of inhibitory substances in pachyman. Fig. 7.1 shows the media isolating the enzyme-producing microor-ganisms described above.
7.2 Alkaline Proteases
Alkaline proteases are not produced only by alkaliphilic microorganisms; two of the most widely studied enzymes are subtilisins BPN’ and Carlsberg produced by neutrophilic strains of Bacillus subtilis. They are not inhibited by metal chelators or thiol reagents; the N-terminal amino acid of both enzymes was identified as alanine and both were inhibited by a serine agent such as diisopropylfluorophosphate (DFP). Several alkaliphilic Streptomyces strains and Bacillus strains produce alkaline proteases capable of hydrolyzing keratinous proteins such as wool, hair, feather and silk under alkaline conditions (in 0.1N NaOH).
7.2.1 Alkaline Protease of Alkaliphilic Bacillus Strains
In 1971 Horikoshi (1971a) reported the production of an extracellular alkaline serine protease from alkaliphilk Bacillus sp. No. 221. This strain, isolated from soil, was found to produce large amounts of alkaline protease which differed from the subtilisin group. The enzyme was purified by DEAE and CM cellulose column chromatography and crystallized from ammonium sulfate solution. The optimum pH of the enzyme was 11.5 with 75% of the activity maintained at pH 13.0 (Fig. 7.2). The enzyme was completely inhibited by DFP or 6 M urea, but not by EDTA or p-chloromercuribenzoate. The molecular weight of the enzyme was 30,000, which is slightly higher than those of other reported alkaline proteases. Calcium ions affected both activity and stability of the enzyme. ...
Table of contents
- Cover
- Half Title
- Title Page
- Copyright Page
- Dedication
- Preface
- Table of Contents
- Introduction
- Isolation, Distribution and Taxonomy of Alkaliphilic Microorganisms
- Cell Structure
- Physiology
- Molecular Biology
- Alkaliphily
- Extracellular Enzymes
- Applications of Alkaliphiles
- References
- Addendum
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Alkaliphiles by Koki Horikoshi,Horikoshi in PDF and/or ePUB format, as well as other popular books in Biological Sciences & Biochemistry in Medicine. We have over 1.5 million books available in our catalogue for you to explore.