
eBook - ePub
Fungal Biotechnology in Agricultural, Food, and Environmental Applications
Dilip K. Arora, Dilip K. Arora
This is a test
Compartir libro
- 524 páginas
- English
- ePUB (apto para móviles)
- Disponible en iOS y Android
eBook - ePub
Fungal Biotechnology in Agricultural, Food, and Environmental Applications
Dilip K. Arora, Dilip K. Arora
Detalles del libro
Vista previa del libro
Índice
Citas
Información del libro
Contributions from 80 world-renowned authorities representing a broad international background lend Fungal Biotechnology in Agricultural, Food, and Environmental Applications first-class information on the biotechnological potential of entomopathogenic fungi and ergot alkaloids, applications of Trichoderma in disease control, and the d
Preguntas frecuentes
¿Cómo cancelo mi suscripción?
¿Cómo descargo los libros?
Por el momento, todos nuestros libros ePub adaptables a dispositivos móviles se pueden descargar a través de la aplicación. La mayor parte de nuestros PDF también se puede descargar y ya estamos trabajando para que el resto también sea descargable. Obtén más información aquí.
¿En qué se diferencian los planes de precios?
Ambos planes te permiten acceder por completo a la biblioteca y a todas las funciones de Perlego. Las únicas diferencias son el precio y el período de suscripción: con el plan anual ahorrarás en torno a un 30 % en comparación con 12 meses de un plan mensual.
¿Qué es Perlego?
Somos un servicio de suscripción de libros de texto en línea que te permite acceder a toda una biblioteca en línea por menos de lo que cuesta un libro al mes. Con más de un millón de libros sobre más de 1000 categorías, ¡tenemos todo lo que necesitas! Obtén más información aquí.
¿Perlego ofrece la función de texto a voz?
Busca el símbolo de lectura en voz alta en tu próximo libro para ver si puedes escucharlo. La herramienta de lectura en voz alta lee el texto en voz alta por ti, resaltando el texto a medida que se lee. Puedes pausarla, acelerarla y ralentizarla. Obtén más información aquí.
¿Es Fungal Biotechnology in Agricultural, Food, and Environmental Applications un PDF/ePUB en línea?
Sí, puedes acceder a Fungal Biotechnology in Agricultural, Food, and Environmental Applications de Dilip K. Arora, Dilip K. Arora en formato PDF o ePUB, así como a otros libros populares de Biological Sciences y Botany. Tenemos más de un millón de libros disponibles en nuestro catálogo para que explores.
Información
1
Biotechnological Approaches in Plant Protection: Achievements, New Initiatives, and Prospects
Sally Ann Leong U.S. Department of Agriculture-Agricultural Research Service, and University of Wisconsin, Madison, Wisconsin, USA
1 CHALLENGES FOR FOOD SECURITY IN THIS CENTURY AND BEYOND
Today we face many critical issues in agriculture: (a) an exponentially growing human population; (b) recurrent famine; (c) the destruction of natural landscapes such as tropical rain forests to extend agriculture to previously unused lands; (d) the exodus of human civilization from rural communities to cities; (e) the destruction of environmental quality resulting from exposure to agrochemicals, erosion of soils and salinization of soils as well as exhaustion and contamination of fresh water resources; (f) the loss of biodiversity through monocropping and the destruction of natural habitats; (g) the reliance of agricultural production, transport, and storage systems on fossil fuel; (h) the acquisition and concentration of agricultural wealth by multinational corporations; and (i) an issuant lack of knowledge by a growing proportion of human civilization on how to cultivate, prepare, and preserve food. The United Nations Food and Agriculture Organization predicts that agricultural productivity in the world will be able to sustain the growing human population by 2030 but hundreds of millions of people in developing countries will remain hungry and environmental problems caused by agriculture will remain serious (FAO 2002). By 2025,83% of the expected global population of 8.5.2 billion will be in the developing world (United Nations 2002). The social consequences are obvious. Food is a basic human need and right. How can we sustain the food needs of the earth’s biotic community in the 21st century and beyond while preserving environmental quality and the diversity and quality of life on earth (Time, August 26, 2002)? What solutions can biotechnology provide to address these problems (Khush and Bar 2001)?
In the last century, the Green Revolution addressed the food needs of the human population through the development of high yielding and early maturing varieties that performed under favorable conditions of nutrition and moisture (Khush 2001). Prior to this time, increased production was dependent on expansion of land area for crop production. In recent years, yield gain through breeding has not kept pace with population growth (Serageldin 1999). Furthermore, genotype is not the only factor limiting productivity. Abiotic and biotic stress factors also contribute to losses in yield both pre and postharvest. For example, in Asia these technical constraints on rice production may reduce production by 23% (Evanson et al. 1996). Socioeconomic constraints also contribute to practices that affect yield. Ninety percent of the world’s rice is grown in Asia on small farms with limited resources. Thus, decisions are made based on economics rather than achieving technically optimum yields. Despite much research on the application of biotechnology to solve these technical production constraints, biotechnology has had limited impact to date on rice production in Asia (Houssain 1997). This has been due in part to the reluctance to adopt the use of genetically modified organisms (GMOs) in most countries of Asia. China is the only Asian country that has embraced biotechnology on any notable scale as a solution to technical production constraints in agriculture (Huang et al. 2002; Pray et al. 2002). For example, over four million smallholders have been able to increase yield and reduce pesticide costs and adverse health effects of applying pesticides by using transgenic Bt-cotton (Pray et al. 2002). Another constraint relates to the complexities of ownership rights that can delay and discourage the scientific application of biotechnology discoveries and their transfer to the market place (Kowalski et al. 2002). Finally, public concern about the safety of consuming GMOs has left tons of food aid containing transgenic corn untouched in Zimbabwe and Zambia despite the prediction that 13 million people are now at risk of famine due to severe drought in Southern Africa (Paarlberg 2002). Organic growers worldwide have rejected GMOs as an allied technology to disease and pest management. Late blight in potato is still very difficult to control in these farming systems (Mader et al. 2002).
Biotechnology as applied to plant protection against fungal pathogens has seen three phases of development over the last 20 years: (a) the application of molecular markers to marker-assisted breeding and the map-based cloning of genes associated with disease resistance and the plant defense response as well as to study fungal pathogenesis and host recognition; (b) the development of routine methods for stable and transient transformation of plants and fungi with foreign genes; and (c) the application of New Biology approaches to study plant growth and development and mechanisms of plant response to abiotic and biotic stresses and fungal pathogenesis. The purpose of this short review is to provide a critical analysis of these recent biotechnological approaches to plant protection with emphasis on fungal pathogens.
2 TESTED STRATEGIES
2.1 Marker-Assisted Breeding and Map-Based Cloning of Genes
2.1.1 Molecular Maps
With the advent of recombinant DNA technology came the application of cloned DNAs as probes to genomic DNA of the source organism and the revelation that different alleles could be detected between individuals based on restriction fragment length polymorphisms (Helentjaris et al. 1985) (Figure 1). Genetic maps based on these and other types of molecular markers (Table 1) were developed for many organisms including crop plants such as rice (McCouch et al. 1988; http://rgp.dna.affrc.go.jp/publicdata/geneticmap2000/index.html), lettuce (Kesseli et al. 1994), tomato (Tanksley et al. 1992), alfalfa (Brouwer and Osborn 1999), and Brassica spp. (Kole et al. 2002), among others. Likewise molecular maps were developed for key fungal pathogens such as Magnaporthe grisea (Farman and Leong 1995; Nitta et al. 1997; Skinner et al. 1993; Sweigard et al. 1993), Phytophtora infestans (van der Lee 2001), and Leptosphaeria maculans (Pongam et al. 1988). These studies also began to reveal the complexities of these genomes in terms of repeated DNAs, their function as transposable elements (Goff et al. 2002; Hamer et al. 1989; Kachroo et al. 1997), their distribution within the genome (Goff et al. 2002; McCouch et al. 1988; Nitta et al. 1997; Yu et al. 2002), and their role in genome evolution and host recognition (Farman 2002; Farman et al. 2002; Kang et al. 2001; Song et al. 1997; 1998). Comparative maps were generated in plants by mapping markers across genera and showed considerable synteny within families of plants (Ahn and Tanksley 1993; Bennetzen and Freeling 1993; Chen et al. 1997; Dunford et al. 1995; Gale and Devos 1998; Hulbert et al. 1990; Saghai Maroof et al. 1996; Tanksley et al. 1992). The mapping of phenotypic markers, both native and induced by mutation, followed closely behind and yielded precise information on the chromosomal location of genes important to plant defense (Ronald et al. 1992; Wang et al. 1995) and fungal host specificity (Dioh et al. 2000; Smith and Leong 1994; Sweigard et al. 1993) and led to their cloning by chromosome walking (Cao et al. 1997; Farman and Leong 1998; Martin et al. 1993; Orbach et al. 2000; Song et al. 1995; Sweigard et al. 1995). The cloning of a plethora of disease resistance genes from many plant species has shown that they belong to a small number of structural classes (Brueggeman et al. 2002; Chauhan and Leong 2002; Dangl and Jones 2001; Meyers et al. 1999; Xiao et al. 2001) (Figure 2). By contrast, the predicted structures of fungal cultivar specificity genes are quite diverse (Bohnert et al. 2001; De Wit and Joosten 1999; Orbach et al. 2000; Sweigard et al. 1995).
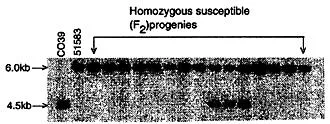

Figure 1 Cosegregation of a RFLP marker (R-23 16) with Pi-CO39(t) locus in homozygous F2 susceptible progenies. Genomic DNA of CO39 (R, resistant), 51583 (S, susceptible) and F2 progenies was digested with Dral, blotted and probed with R-2316. Recombinant progenies show DNA fragments from both parents. Phosphoimage of Southern blot is shown.
These studies have been complemented by the mapping of candidate genes such as the PR (pathogenesis-related) proteins in plants that were discovered from differential expression of RNA and protein during plant infection (Muthukrishnan et al. 2001) or resistance gene analogs based on the conserved structural features of disease resistance genes (Boyko et al. 2002; Chauhan et al. 2002; Faris et al. 1999; Gebhardt and Valkonen 2001; Huang and Gill 2001; Li et al. 1999; Shen et al. 1998; Speulman et al. 1998). This analysis has been particularly well advanced in wheat and its relative Aegilops tauschii. Resistance and defense response genes in A. tauschii are localized in clusters primarily in while in Chinese spring wheat defense response genes are localized in clusters and/or at distal regions of chromosomes (Li et al. 1999). In many cases, these genes or gene homologs have been correlated with loci that affect quantitative or distal/telomeric regions of the genome (Boyko et al. 2002) single gene resistance in the respective plants. For example, QTLs with large effects in wheat were shown to contain RGAs or clusters of defense response genes such as catalase, chitinase, thaumatins, and an ion channel regulator (Faris et al. 1999). Similar results are emerging in the genomes of potato (Gebhardt and Valkonen 2001), Arabidopsis (Speulman et al. 1998) and pepper (Pflieger et al. 2001).
Table 1 Molecular markers used in mapping of traits
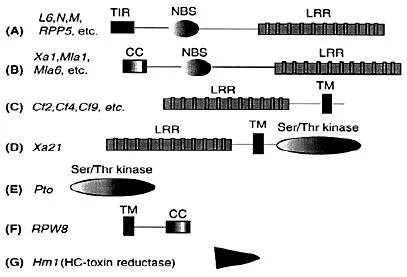
Figure 2 Different classes (A-G) of plant disease resistance genes [reviewed in Chauhan and Leong (2002)): Genes in classes A and B are cytoplasmic proteins differing only in their N-terminal domains; Class C genes encode putative transmembrane molecules with an extracellular LRR domain; Xi21 is a transmembrane protein with an extracellular LRR domain; Pto is a cytoplasmic Ser/Thr kinase; RPW8 contains a putative Nterminal TM domain and a CC domain; Hm1 is a unique enzyme that detoxifies a fungal toxin; Abbreviations for domains: TIR, Drosophila Toll/Human Interleukinlreceptor; CC, Coiled-coil; NBS, Nucleotide binding site; LRR, Leucine-rich repeat; TM, Transmembrane; Ser/Thr, Serine/threonine kinase.
Efforts are underway to functionally characterize 179 NBS-LRR-encoding genes that may encode disease resistance genes in the Arabidopsis genome (Figure 2). These have been organized into subclasses and their distribution mapped to the chromosomal sequence (Michelmore 2002; www.nibh-rs.ucdavis.edu). A publicly available, draft ordered sequence of the rice genome is anticipated by the end of 2002 (http://rgp.dna.affrc.go.jp/cgi-bin/statusdb/seqcollab.pl) and will allow comparisons to be done across syntenic regions of grass genomes (http://www.gramene.org!/). The unordered draft sequence of rice varieties Nipponbare and 93–11 has revealed numerous NBS-LRR-containing sequences as well as sequences potentially encoding other minor classes of R genes and Arabidopsis genes known to control defense response signal transduction (Goff et al. 2002; Yu 2002). Preliminary studies based on conservation of RGAs in comparative maps of the grasses have shown evidence for some conservation but also redistribution of this class of genes among the grasses (Leister et al. 1998). Likewise a detailed comparison of a syntenic region between barley and rice did not reveal any candidate resistance genes in rice that could be the ortholog of Rpgl in barley (Han et al. 1999; Kilian et al. 1997).
2.1.2 Differential cDNA-AFLP Screens
Differential cDNA-APLP screening has been done to isolate hypersensitive response (HR)-specific genes to the Cladesporium fulvum elicitor Avr4 in tomato and has led to the isolation of a previously known and corresponding disease resistance gene cluster Cf- 4 as well as numerous new candidate genes involved in the HR response (De Wit et al. 2002; Takken et al. 2001). This method is a robust and inexpensive way to identify differentially expressed genes involving the digestion of cDNAs with two different restriction enzymes and the amplification of the resulting products after ligation to adapters for these enzymes. The sizes of the resulting amplicons are measured by gel electrophoresis and resulting fragments can be excised and sequenced. Comparison of this method with differential display has shown the cDNA-AFLP method to be superior (Jones and Harrower 1998). Using the cDNA-AFLP technique, Durrant et al. (2000) found a strong coincidence between the expression of genes involved in race-specific resistance and the wound response in tobacco cell cultures. Collectively, these candidate genes will provide additional markers for studies of disease resistance traits in the potato and tomato genomes. Infection-specific cDNA-AFLPs have also been identified in Arabidopsis thaliana inoculated with Peronospora parasitica (van der Biezen et al. 2000). Interestingly, most fragments were derived from the fungal pathogen showing the power of this method to study genes expressed in the pathogen, which in many cases may represent a minor component of the mass of the tissue studied.
Previous genetic studies by Valent et al. (1991) have shown that infection of some grass hosts by M. grisea is a quantitatively inherited trait. The cDNA-APLP approach would allow for the identification a unique set of cDNA-AFLPs in each progeny showing varying degrees of pathogenicity and in some cases segregating with pathogenicity. This approach has been recently used to create genome-wide transcription maps of Arabidopsis and potato and study inheritance of the cDNA-AFLPs in segregating populations (Brugmans et al. 2002). Thus phenotypes can be directly associated with molecular genotypes and candidate gene fragments can be excised from gels for further analysis. We are using this approach in an attempt to identify major and minor genes controlling resistance to blast and drought tolerance in Eleusine coracana.
Computational methods for relating the size of the AFLP restriction fragment products to the predicted restriction fragment products from sequenced cDNA libraries have been developed and used to identify putative, infection stagespecific, pathogenicity factors from the plant pathogenic nematode Globoderu rostochiensis without need for sequencing the gel fragments (Qin et al. 2001). For those organisms having fully sequenced, annotated full length cDNA libraries such as Arabidopsis (Seki et al. 2002), this approach provides for the rapid functional classification of the cDNA-AFLPs.
2.1.3 Identification of QTL-Associated Genes
Very few QTL studies in plants have led to the cloning of a single gene within a QTL that is responsible for the variation seen [reviewed in Buckler and Thornsberry (2002)]. These few examples represent QTLs that had major effects on variation. Buckler and Thornsberry (2002) have proposed that association approaches should be also considered to provide improved resolution and to reduce the time of analysis as mapping populations are not needed since natural variation in a population is investigated instead. The resolution of association that can be obtained depends on the linkage disequilibrium (LD) structure of the population of organism being studied and some insight on candidate gene(s) to target. Studies on LD structure have shown that inbreeding plants such as Arabidopsis have large LD structures on the order of 250 kb or 1 cM (Nordborg et al. 2002) while outbreeding plants like maize have very small LD structures in the order of kbs (Buckler and Thornsberry 2002). Using this approach, Thornsberry et al. (2001) were able to associate polymorphisms found in the Dwarf8 gene of maize with variation seen in flowering time.
In Saccharomyces cereviseae whose entire genome sequence is known, a QTL for high temperature growth (Htg) commonly found in clinical isolates was rigorously analyzed to identify the responsible genes (Steinmetz et al. 2002). Using reciprocal hemizygosity tests, involving selective gene disruption of candidate genes in both parental genomes and then forming diploid hybrids among these strains, three genes were found to contribute to the phenotype in the QTL interval. However, the alleles of two genes came from one parent strain while that of the third came from the other parent. By contrast attempts to employ natural sequence variation or mRNA expression levels determined from several natural isolates of yeast did not provide a clue to which gene(s) in the interval contributed to the phenotype. Thus employing LD and association by decent to accelerate gene identification in an interval may not always succeed and genetic studies will be required to study inheritance and create reciprocal hemizygotes in targeted regions of the genome. The use of allele-specific gene silencing methods (see below) may allow this to be done in the F1 generation of plants while targeted gene disruption methods can be used in fungi such as Ustilago maydis that have a stable diploid phase and facile gene knockout system. Transformation of haploid fungi with wild type and disrupte...