![]()
1
Introduction to forensic genetics
The development and application of genetics has revolutionized forensic science. In 1984, the analysis of polymorphic regions of DNA produced what was termed ‘a DNA fingerprint’ [1]. The following year, at the request of the United Kingdom Home Office, DNA profiling was successfully applied to casework when it was used to resolve an immigration dispute [2]. In 1986, DNA evidence was used for the first time in a criminal case involving the murder of two young women in Leicestershire, UK: DNA analysis exonerated one individual who had confessed to one of the murders, and following a mass screen of approximately 5000 individuals, identified Colin Pitchfork as the murderer. He was convicted in January 1988 [3].1
Following on from early success in both civil and criminal cases, the use of genetics was rapidly adopted by the forensic community and now plays an important role worldwide in both the investigation of crime and in relationship testing. The scope and scale of DNA analysis in forensic science is set to continue expanding for the foreseeable future.
Forensic genetics
The work of the forensic geneticist will vary widely depending on the laboratory and country that they work in, and can involve the analysis of material recovered from a scene of crime, kinship testing and the identification of human remains. In some cases, it can even be used for the analysis of DNA from plants [6–18]; animals [19–36] including insects [37–60]; and microorganisms [61–67]. The focus of this book is the analysis of biological material that is recovered from the scene of crime – this is central to the work of most forensic laboratories. Kinship testing will be dealt with separately in Chapter 11 and a brief introduction is given to the testing of non-human material in Chapter 14.
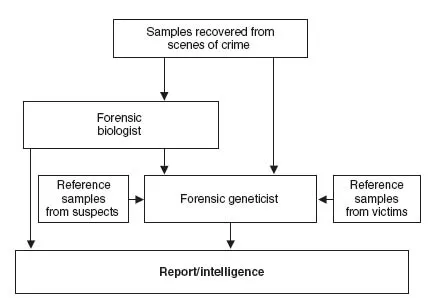
Forensic laboratories receive material that has been recovered from scenes of crime, and reference samples from both suspects and victims. The role of forensic genetics within the investigative process is to compare samples recovered from crime scenes with suspects and possibly victims, resulting in a report that can be presented in court or intelligence that may inform an investigation (Figure 1.1).
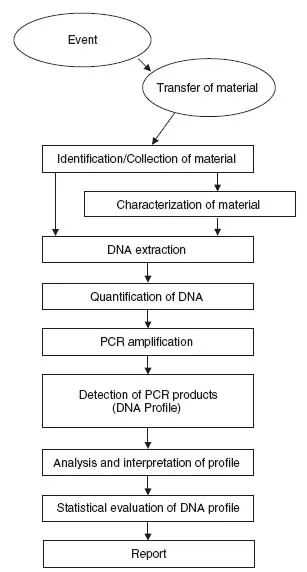
Several stages are involved in the analysis of genetic evidence (Figure 1.2) and each of these is covered in detail in the following chapters. In some organizations one person will be responsible for collecting the evidence, the biological and genetic analysis of samples and ultimately presenting the results to a court of law. However, the trend in many larger organizations is for individuals to be responsible for only a very specific task within the process, such as the extraction of DNA from the evidential material or the analysis and interpretation of DNA profiles that have been generated by other scientists, or just reporting the findings.
A brief history of forensic genetics
In 1900 Karl Landsteiner described the ABO blood grouping system and observed that individuals could be placed into different groups based on their blood type. This was the first step in the development of forensic haemogenetics. Following on from this, numerous blood group markers and soluble blood serum protein markers were characterized and could be analysed in combination to produce highly discriminatory profiles. The serological techniques were a powerful tool but were limited in many forensic cases by the amount of biological material that was required to provide highly discriminating results. Proteins are also prone to degradation on exposure to the environment.
In the 1960s and 1970s, developments in molecular biology, including restriction enzymes, Sanger sequencing [68] and Southern blotting [69], enabled scientists to examine DNA sequences. By 1978, DNA polymorphisms could be detected using Southern blotting [70] and in 1980 the analysis of the first highly polymorphic locus was reported, where the polymorphism was caused by differences in the lengths of the alleles [71].
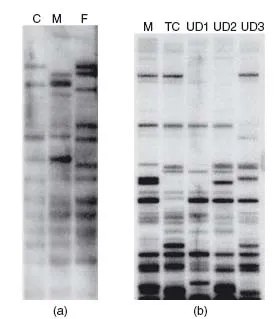
It was not until September 1984 that Alec Jeffreys realized the potential forensic application of the minisatellite loci [1, 72, 73]. The technique developed by Jeffreys entailed extracting DNA and cutting it with a restriction enzyme before carrying out agarose gel electrophoresis, Southern blotting and probe hybridization to detect the polymorphic loci. The end result was a series of black bands on X-ray film, which was called a DNA fingerprint (Figure 1.3).
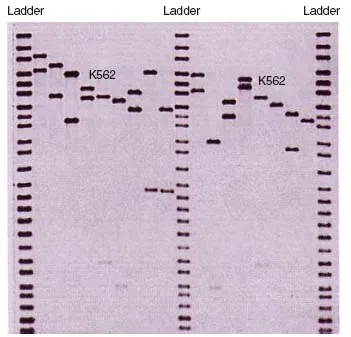
With the first DNA fingerprints the multi-locus probes (MLPs) detected several minisatellite loci simultaneously, leading to the multiple band patterns. While the multi-banded fingerprints were highly informative they were difficult to interpret. New probes were designed that were specific to one locus (single locus probes, SLPs) and therefore produced only one or two bands for each individual [74] (Figure 1.4).
Minisatellite analysis was a powerful tool but suffered from several limitations: a relatively large amount of DNA was required; it would not work with degraded DNA; comparison between laboratories was difficult; and the analysis was time consuming. Even so, the use of minisatellite analysis, using SLPs, was common for several years [75] until it was replaced by polymerase chain reaction (PCR)-based systems.
A critical development in the history of forensic genetics came with the advent of a process that can amplify specific regions of DNA – the PCR (see Chapter 5). The PCR process was conceptualized in 1983 by Kary Mullis, a chemist working for the Cetus Corporation in the USA [76]. The development of PCR has had a profound effect on all aspects of molecular biology including forensic genetics, and in recognition of the significance of the development of the technique Kary Mullis was awarded the Nobel Prize for Chemistry in 1993.
The PCR increases the sensitivity of DNA analysis to the point where DNA profiles can be generated from just a few cells, reduces the time required to produce a profile, can be used with degraded DNA and allows just about any polymorphism in the genome to be analysed.
The first application of PCR in a forensic case involved the analysis of single nucleotide polymorphisms in the human leukocyte antigen (HLA)-DQα locus (part of the major histocompatibility complex (MHC)) [77] (see Chapter 12). This was soon followed by the analysis of short tandem repeats (STRs), which are currently the most commonly used genetic markers in forensic science (see Chapters 6–8). The rapid development of technology for analysing DNA includes advances in DNA extraction and quantification methodology, the development of commercial PCR-based typing kits and equipment for detecting DNA polymorphisms.
In addition to technical advances, another important part of the development of DNA profiling that has had an impact on the whole field of forensic science is quality assurance. The admissibility of DNA evidence was seriously challenged in the USA in 1989 – People v. Castro [78]; this and subsequent cases in many countries have resulted in increased levels of standardization and quality assurance in forensic genetics and other areas of forensic science. As a result, the accreditation of both laboratories and individuals is an increasingly important issue in forensic science. The combination of technical advances, high levels of standardization and quality assurance have led to forensic DNA analysis being recognized as a robust and reliable forensic tool.
References
1. Jeffreys, A.J. Wilson, V. and Thein, S.L. (1985) Individual-specific fingerprints of humanDNA. Nature, 316, 76–79.
2. Jeffreys, A.J., Brookfield, J.F.Y. and Semenoff, R. (1985) Positive identification of an immigration test-case using human DNA fingerprints. Nature, 317, 818–819.
3. Seton, C. (1988) Life for Sex Killer who Sent Decoy to Take Genetic Test. The Times(London).
4. McCarthy, M. (1987) Rapist in Genetic Fingerprint Case Jailed for 8 Years. The Times(London).
5. Andrews, T.L. (1987) State v. Andrews. 9th Cir. Ct, Orange Co., Fla, Div 15.
6. Kress, W.J., Wurdack, K.J., Zimmer, E.A., Weigt, L.A. and Janzen, D.H. (2005) Use of DNA barcodes to identify flowering plants. Proceedings of the National Academy of Sciences of theUnited States of America, 102, 8369–8374.
7. Linacre, A. and Thorpe, J. (1998) Detection and identification of cannabis by DNA. ForensicScience International, 91, 71–76.
8. Bless, C., Palmeter, H. and Wallace, M.M. (2006) Identification of Acer rubrum using amplifiedfragment length polymorphism. Journal of Forensic Sciences, 51, 31–38.
9. Cowan, R.S., Chase, M.W., Kress, W.J. and Savolainen, V. (2006) 300,000 species to identify:problems, progress, and prospects in DNA barcoding of land plants. Taxon, 55, 611–616.
10. Craft, K.J., Owens, J.D. and Ashley, M.V. (2007) Application of plant DNA markers inforensic botany: genetic comparison of Quercus evidence leaves to crime scene trees usingmicrosatellites. Forensic Science International, 165, 64–70.
11. Datwyler, S.L. and Weiblen, G.D. (2006) Genetic variation in hemp and marijuana (Cannabissativa L.) according to amplified fragment length polymorphisms. Journal of Forensic Sciences, 51, 371–375.
12. Ferri, G., Alu, M., Corradini, B. and Beduschi, G. (2009) Forensic botany: species identificationof botanical trace evidence using a multigene barcoding approach. International Journal ofLegal Medicine, 123, 395–401.
13. Finkeldey, R., Leinemann, L. and Gailing, O. (2010) Molecular genetic tools to infer the originof forest plants and wood. Applied Microbiology and Biotechnology, 85, 1251–1258.
14. Howard, C., Gilmore, S., Robertson, J. and Peakall, R. (2008) Developmental validation of aCannabis sativa STIR multiplex system for forensic analysis. Journal of Forensic Sciences,53, 1061–1067.
15. Howard, C., Gilmore, S., Robertson, J. and Peakall, R. (2009) A Cannabis sativa STR genotypedatabase for Australian seizures: forensic applications and limitations. Journal of ForensicSciences, 54, 556–563.
16. Kojoma, M., Seki, H., Yoshida, S. and Muranaka, T. (2006) DNA polymorphisms in thetetrahydrocannabinolic acid (THCA) synthase gene in “drug-type” and “fiber-type” Cannabissativa L. Forensic Science International, 159, 132–140.
17. Kress, W.J., Erickson, D.L., Jones, F.A., Swenson, N.G., Perez, R., Sanjur, O. et al. (2009)Plant DNA barcodes and a community phylogeny of a tropical forest dynamics plot inPanama. Proceedings of the National Academy of Sciences of the United States of America,106, 18621–18626.
18. Tsai, L.C., Yu, Y.C., Hsieh, H.M., Wang, J.C., Linacre, A. and Lee, J.C.I. (2006) Speciesidentification using sequences of the trnL intron and the trnL-trnF IGS of chloroplast genomeamong popular plants in Taiwan. Forensic Science International, 164, 193–200.
19. Hebert, P.D.N., Ratnasingham, S., deWaard, J.R. (2003) Barcoding animal life: cytochrome coxidase subunit 1 divergences among closely related species. Proceedings of the Royal Societyof London Series B: Biological Sciences, 270, S96–S99.
20. Parson, W., Pegoraro, K., Niederstatter, H., Foger, M. and Steinlechner, M. (2000) Speciesidentification by means of the cytochrome b gene. International Journal of Legal Medicine,114, 23–28.
21. Eichmann, C. and Parson, W. (2007) Molecular characterization of the canine mitochondrialDNA control region for forensic applications. International Journal of Legal Medicine, 121,411–416.
22. Fernandes, C.A., Ginja, C., Pereira, I., Tenreiro, R., Bruford, M.W. and Santos-Reis, M. (2008)Species-specific mitochondrial DNA markers for identification of non-invasive samples fromsympatric carnivores in the Iberian Peninsula. Conservation Genetics, 9, 681–690.
23. Halverson, J.L. and Basten, C. (2005) Forensic DNA identification of animal-derived traceevidence: tools for linking victims and suspects. Croatian Medical Journal, 46, 598–605.
24. Hellmann, A.P., Rohleder, U., Eichmann, C., Pfeiffer, I., Parson, W. and Schleenbecker, U.(2006) A proposal for standardization in forensic canine DNA typing: allele nomenclature ofsix canine-specific STR loci. Journal of Forensic Sciences, 51, 274–281.
25. Hsieh, H.M., Huang, L.H., Tsaim L.C., Liu, C.L., Kuo, Y.C., Hsiao, C.T. etal. (2006) Speciesidentification of Kachuga tecta using the cytochrome b gene. Journal of Forensic Sciences,51, 52–56.
26. Imaizumi, K., Akutsu, T., Miyasaka, S. and Yoshino, M. (2007) Development of speciesidentification tests targeting the 16S ribosomal RNA coding region in mitochondrial DNA.International Journal of Legal Medicine, 121, 184–191.
27. Lee, J.C.I., Tsai, L.C., Yang, C.Y., Liu, C.L., Huang, L.H., Linacre, A. et al. (2006) DNAprofiling of shahtoosh. Electrophoresis, 27, 3359–3362.
28. Magnussen, J.E., Pikitch, E.K., Clarke, S.C., Nicholson, C., Hoelz...