
eBook - ePub
Quantitative Structure-Activity Relationship (QSAR) Models of Mutagens and Carcinogens
- 304 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Quantitative Structure-Activity Relationship (QSAR) Models of Mutagens and Carcinogens
About this book
Applied with success in a number of areas, QSAR studies have become particularly popular in the rational design of drugs and pesticides. Much has been published on the principles of QSAR in this area, but not on their application s to toxic chemicals.
This book provides the first comprehensive, interdisciplinary presentation of QSAR studies on
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Topic
MedicineSubtopic
Biochemistry in Medicine1 General Introduction to QSAR
CONTENTS
1.1 Introduction
1.2 Some Basic Principles
1.3 Free–Wilson Analysis
1.4 Hansch Analysis
1.4.1 Basic Assumptions
1.4.2 Parameters
1.4.2.1 Electronic Parameters
1.4.2.2 Hydrophobic Parameters
1.4.2.3 Steric Parameters
1.4.2.4 Indicator Variables
1.4.3 Building and Evaluating Hansch Equations
1.5 Some Multivariate Methods
1.5.1 Principal Components and PLS
1.5.2 Three-Dimensional QSAR
1.5.3 Classification Methods
1.6 Some Other QSAR-Related Methods
1.7 Concluding Remarks
References
1.1 INTRODUCTION
Classical chemometric QSAR methods for the analysis of quantitative structure– activity relationships (QSARs) are sometimes regarded to be out of fashion when compared with the rapid development of molecular modeling, structure-based design, and protein crystallography. In addition, an equation is more difficult to understand than a colored three-dimensional picture generated by computer graphics. However, classical QSAR methods still play an important role and will continue to be a useful tool in modern drug design.1–3 They have contributed greatly to the development of science in medicinal chemistry (QSAR “know how”), and thousands of documented QSARs and success stories of QSAR predictions and QSAR-guided drug design attest to their versatility. In particular, the quantitative description of pharmacokinetic processes remains the domain of classical QSAR techniques. This aspect and QSAR-based concepts such as “drug likeness” are gaining in importance in connection with high throughput screening (HTS) for hit to lead decisions in order to avoid the selection of compounds with unfavorable adsorption/distribution/ metabolism/excretion (ADME) properties. Another important issue is the design of safe and selective compounds and a better understanding of toxic, carcinogenic, or mutagenic effects.
This chapter presents a condensed introduction to the most important classical QSAR methods with the main emphasis on Free–Wilson and Hansch analyses. Only references absolutely essential for the understanding of the text will be presented with no attempt for completeness in the sense of a review. For a follow-up, the reader is referred to a number of monographs2–21 on various aspects of the QSAR field, to the proceedings of the European QSAR conferences (see References 22 to 25 for the last four meetings), and to the journal Quantitative Structure–Activity Relationships, which provides an excellent and exhaustive abstract service.
1.2 SOME BASIC PRINCIPLES
Probably the first general formulation of a quantitative structure–activity relationship was presented by Crum-Brown and Fraser in 1868 who assumed that biological activity is a function of chemical structure (“constitution”):
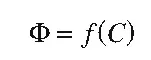
(1.1)
From this general formulation to the development of true QSARs was still a long way to go because it was necessary to define proper measures of F, suitable mathematical formalisms for the function f, and methods to quantitatively describe chemical structure C. Modern QSAR technology started in 1964 with publications by Hansch and Fujita26 and Free and Wilson.27 The first publication led to development of the well-known Hansch analysis, the most widely-used QSAR method also known as the extrathermodynamic or linear free-energy-related approach. The second paper resulted in development of the so-called Free–Wilson analysis, which supplements Hansch analysis and has turned out to be a very useful method for certain types of structural modifications. Both methods use multiple regression analysis as the mathematical method (f in Equation (1.1)) but differ in the description of chemical properties. In Hansch analysis, substituent constants and other physicochemical descriptors are used, while Free–Wilson analysis is based on chemical fragments directly derived from the two-dimensional structure of compounds.
Today, a large variety of mathematical methods is available to express the f in Equation (1.1). To name just a few, the most frequently used methods are multiple regression analysis, principal component and factor analysis, principal component regression analysis, partial least squares (PLS), discriminant analysis and other classification methods, and neuronal nets. The variety of mathematical methods is accompanied by a huge number of chemical descriptors to characterize chemical structure; an impressive encyclopedic guide to such descriptors has been presented by Todeschini and Consonni in their Handbook of Molecular Descriptors.28 Not all of these descriptors have proven to be useful. Broadly speaking, they may be categorized as experimental quantities, such as log P, pKa (these quantities can also be computed; see below), and spectroscopic data; substituent constants (electronic, hydrophobic, and steric); parameters derived from molecular modeling and quantum chemical computations; graph theoretical indices; and variables describing the presence or the number of occurrences of certain substructures.
Typical measures of biological activity are the molar concentration C of a compound producing a certain effect derived from a dose–response curve (e.g., ED50 or IC50); binding, association, or inhibition constants; and rate constants. In order to obtain larger values for more active compounds, reciprocal values are usually considered for dissociation constants and the molar-concentration-based quantities. Based on thermodynamic or kinetic reasoning, such parameters can be turned into free-energy-related quantities by logarithmic transformation, which is required for the formalism of Hansch analysis (for a detailed discussion, see Franke7). Thus, typical expressions for Φ in Equation (1.1) are pC = –log C = log 1/C (examples: pED50 or pIC50), log K (where K is a binding, inhibition, or rate constant), and log 1/Kd (where Kd is a dissociation constant). By convention, the logarithmic transformation of biological measurement is used not only in Hansch analysis (or other methods based on linear free energy relationships) but in all QSAR approaches applied to quantitative (continuous) biological measurements. One of the reasons is that the results are better comparable. Sometimes, biological measurements result in %effect data measured at a single dose. Strictly speaking, such data are not suitable for Hansch-type and related QSAR approaches. Experience has shown, however, that such data can still lead to meaningful QSARs after logarithmic transformation, provided that the entire range from a few percent values to values close to 100% is covered. A good alternative for such values is a logit transformation according to:
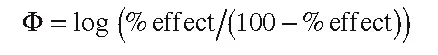
(1.2)
Another alternative is to translate %effect data into a classification scheme that can then be analyzed by classification methods. Such methods are also necessary if biological measurements only allow a scoring of biological potency. In the following text, the logarithmically transformed activity values will be designated as log BR (BR = biological response).
1.3 FREE–WILSON ANALYSIS
The Free–Wilson analysis can be applied to series of compounds where the compounds consist of a common (constant) parent structure and variable fragments (usually substituents) (see Figure 1.1). The basic assumptions of Free–Wilson analysis are:...
Table of contents
- COVER PAGE
- TITLE PAGE
- COPYRIGHT PAGE
- PREFACE
- THE EDITOR
- CONTRIBUTORS
- ABSTRACT
- 1. GENERAL INTRODUCTION TO QSAR
- 2. MECHANISMS OF ACTION OF CHEMICAL CARCINOGENS AND THEIR ROLE IN STRUCTURE–ACTIVITY RELATIONSHIPS (SAR) ANALYSIS AND RISK ASSESSMENT
- 3. QSARs FOR INDIVIDUAL CLASSES OF CHEMICAL MUTAGENS AND CARCINOGENS
- 4. QSARs FOR THE MUTAGENICITY AND CARCINOGENICITY OF THE AROMATIC AMINES
- 5. PUBLIC SOURCES OF MUTAGENICITY AND CARCINOGENICITY DATA: USE IN STRUCTURE–ACTIVITY RELATIONSHIP MODELS
- 6. SAR IN THE ASSESSMENT OF CARCINOGENESIS: THE MULTICASE APPROACH
- 7. PREDICTING MUTAGENICITY OF CONGENERIC AND DIVERSE SETS OF CHEMICALS USING COMPUTED MOLECULAR DESCRIPTORS: A HIERARCHICAL APPROACH
- 8. QUANTITATIVE STRUCTURE–ACTIVITY RELATIONSHIPS FOR ACUTE AQUATIC TOXICITY: THE ROLE OF MECHANISM OF TOXIC ACTION IN SUCCESSFUL MODELING
- 9. SARs AND QSARs OF MUTAGENS AND CARCINOGENS: UNDERSTANDING ACTION MECHANISMS AND IMPROVING RISK ASSESSMENT
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Quantitative Structure-Activity Relationship (QSAR) Models of Mutagens and Carcinogens by Romualdo Benigni in PDF and/or ePUB format, as well as other popular books in Medicine & Biochemistry in Medicine. We have over 1.5 million books available in our catalogue for you to explore.