
eBook - ePub
RNA Methodologies
Laboratory Guide for Isolation and Characterization
- 876 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
RNA Methodologies
Laboratory Guide for Isolation and Characterization
About this book
RNA Methodologies, Fifth Edition continues its tradition of excellence in providing the most up-to-date ribonucleic acid lab techniques for seasoned scientists and graduate students alike. This edition features new material on the exploding field of microRNA as well as the methods for the profiling of gene expression, both which have changed considerably in recent years. As a leader in the field, Dr. Farrell provides a wealth of knowledge on the topic of RNA while also giving readers helpful hints from his own personal experience in this subject area. Beginning with the most contemporary, RNA Methodologies, Fifth Edition, presents the essential techniques to use when working with RNA for the experienced practitioner while at the same time providing images and examples to aid the beginner in fully understanding this important branch of molecular biology. The next generation of scientists can look to this work as a guide for ensuring high productivity and highly representative data, as well as best practices in troubleshooting laboratory problems when they arise.
- Features new material in miRNA, MIQE guidelines, biomarkers, RNA sequencing, digital PCR and more
- Includes expanded coverage on quantitative PCR techniques, RNAi, bioinformatics, the role of locked nucleic acids, aptamer biology, PCR arrays, and other modern technologies
- Presents comprehensive, cutting-edge information covering all aspects of working with RNA
- Builds from basic information on RNA techniques to in-depth protocols to guidance on how to modify and adjust each step of a particular application
- Presents multiple avenues for addressing the same experimental goals
Trusted by 375,005 students
Access to over 1 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Chapter 1
RNA and the Cellular Biochemistry Revisited
Abstract
This chapter focuses on the forms and functions of ribonucleic acid (RNA) and its transcription. RNA is a long, unbranched polymer of ribonucleoside monophosphate moieties joined together by phosphodiester linkages, and both eukaryotic and prokaryotic RNAs are essentially single-stranded molecules. RNA molecules are produced by the process of transcription, the process by which a single-stranded RNA molecule is synthesized from a specific chromosome locus. There are important organizational differences in associated with genes and the ensuing RNA molecule that results from the transcription when comparing prokaryotic and eukaryotic organisms. The synthesis of RNA is mediated by the activity of enzymes known as RNA polymerases. These transcripts observed within a cell are traditionally classified as ribosomal RNA (rRNA), transfer RNA (tRNA), heterogeneous nuclear RNA (hnRNA), or messenger RNA (mRNA). Important new classes of RNA have been discovered, including microRNA (miRNA), circular RNA (circRNA), and noncoding RNA (ncRNA). The mRNA subpopulation drives the phenotype of the cell, although it is the least abundant of all transcript types and the expression of which is tightly coupled to the expression of other regulatory transcripts.
Keywords
Transcriptome; gene expression; miRNA; circRNA; operon; splicing; ncRNA; RNA polymerase; transcriptional regulation; posttranscriptional regulation; polynucleotide; transcription
Why Study RNA?
All cell and tissue functions are ultimately governed by gene expression. Consequently, the reasons for electing to study the modulation of RNA levels as at least one parameter of the cellular biochemistry may be as diverse as the intracellular RNA population itself. Generally speaking, the characterization of RNA is almost always related to transcription, i.e., gene expression questions being asked in the context of a particular scientific inquiry, and most often revolves around measuring the dynamic abundance level of one or several transcripts.
The goals in any experimental design involving RNA generally revolve around one or more fundamental themes, including but not limited to the following:
1. Measurement of the steady-state abundance of cellular transcripts. Steady-state RNA refers to the net accumulation of transcription products in the cell, or in a subcellular compartment such as the nucleus or the cytoplasm. It is the combined result of RNA synthesis, stability, and degradation. This is the most common reason why RNA is isolated from cells and tissues. Analysis may focus on one transcript, a few transcripts, or all transcripts simultaneously; this latter approach is commonly known as global analysis of gene expression or whole transcriptome profiling. Given the ease with which RNA can be purified from biological sources, the use of various sensitive, contemporary approaches is widespread for generating quantitative and qualitative profiles of RNA populations using any of a variety of laboratory techniques.
2. Synthesis of complementary DNA (cDNA). Unstable, single-stranded messenger RNA (mRNA) can serve as the template for the in vitro synthesis of very stable single- or double-stranded cDNA molecules. This is the first step for subsequent amplification by the polymerase chain reaction (PCR), often for some “quantitative purpose,” for transcript mapping purposes, for direct ligation into a vector for sequencing or for expression of the encoded protein, for the physical separation of two or more cDNA species, or for the older strategy of synthesizing an entire cDNA library (older literature occasionally refers to a cDNA library as a “clone bank.”) which can be propagated for long-term storage and analysis. In any event, the construction of cDNA is the creation of a permanent biochemical record of the cell at the moment of cellular disruption. Historically, the synthesis of highly representative cDNA is one of the most important methodologies in the molecular biology laboratory and, in some hands, remains a significant challenge.
3. Detection of viruses which harbor an RNA genome. This proceeds via the synthesis of cDNA, as described above, followed by PCR or another cDNA amplification method.
4. Identification of the transcription start site (TSS). Historically, mapping of RNA molecules, including the 5′ end, the 3′ end, and the size and location of introns, was accomplished via nuclease protection assay, as described in Chapter 18, Quantification of Specific mRNAs by Nuclease Protection. Now, however, transcript mapping is now almost always performed by some variant of 5′- or 3′-rapid amplification of cDNA ends (RACE; see Chapter 8: RT-PCR: A Science and an Art Form). As it is well known that a single genetic locus often has the potential to produce multiple RNAs, each with a different TSS and often in a tissue-specific manner, TSS mapping is an invaluable technique.
5. Measurement of the rate of transcription of gene sequences or the pathways of RNA processing. This may be deduced, at least in part, by the nuclear run-on assay in which radiolabeled ribonucleotide precursors are incorporated into nascent transcripts in direct proportion to the abundance of each species of RNA being transcribed (see Chapter 19: Analysis of Nuclear RNA). When used in conjunction with other methods that examine steady-state RNA levels, the regulation of genes can often be assigned as transcriptional or due to posttranscriptional events.
6. In vitro translation of purified mRNA. The resulting polypeptide may be further characterized by immunoprecipitation or Western analysis. Cell-free translation represents an older method for the identification of specific transcripts: by providing the raw materials needed to support translation, one is able to demonstrate that a transcript of putative identity is able to support the synthesis of the cognate peptide. For example, this approach could be used to demonstrate that two transcripts from the same genetic locus with alternative TSSs are, in fact, able to direct the synthesis of identical or closely related proteins. In applications such as rational drug design, in vitro translation is helpful because understanding the three-dimensional architecture of a protein, and its wild type or mutated function(s), may suggest novel applications in the area of functional proteomics.
What is RNA?
RNA is a long, unbranched polymer of ribonucleoside monophosphate moieties joined together by phosphodiester linkages. Both eukaryotic and prokaryotic RNAs are single-stranded molecules. The unlinked monomer building blocks of both RNA and DNA are known generically as nucleotides. Each nucleotide consists of three key components: a pentose (five-carbon sugar), at least one phosphate group (nucleotides may contain as many as three phosphate groups), and a nitrogenous base (Fig. 1.1). A nitrogenous base joined to a pentose sugar is known as a nucleoside. When a phosphate group is added, the composite, a phosphate ester of the nucleoside, is known as a nucleotide.
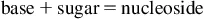
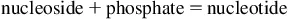

The components of RNA and DNA nucleosides and nucleotides are compared in Table 1.1.
Table 1.1
Comparative Nucleotide Structure
| RNA Nucleotides | DNA Nucleotides | |
| Key Nucleotide Components | Five-carbon ribose | Five-carbon deoxyribose |
| Phosphate group(s) | Phosphate group(s) | |
| Nitrogenous base | Nitrogenous base | |
| Common Nitrogenous Bases | ||
| Purines | Adenine | Adenine |
| Guanine | Guanine | |
| Pyrimidines | Cytosine | Cytosine |
| Uracil | Thymine | |
The key chemical difference between RNA and DNA is the presence the five-carbon sugar ribose, in which a hydroxyl group (–OH) is joined to the 2′ carbon of the ribose sugar, in the case of RNA; the absence of the 2′ OH group in DNA is the underlying basis of the name of the sugar “deoxyribose.” In addition, the base uracil is found in RNA, substituted in DNA by the closely pyrimidine thymine (Chemically, thymine is 5-methyluracil), though it is possible to find deoxynucleotides containing uracil in certain situations. More precisely, RNA is assembled from ribonucleotide precursors and DNA is assembled from deoxyribonucleotide precursors. Hence, RNA is so-named because of the ribose sugar it contains, just as DNA is named from its constituent 2′-deoxyribose sugar. Essential base, nucleoside, and nucleotide nomenclature is summarized in Table 1.2.
Table 1.2
Essential Base, Nucleoside, and Nucleotide Nomenclature
| Base | Nucleoside | Nucleotide Triphosphate Precursors | |
| RNA | Adenine (A) | Adenosine | Adenosine-5′-triphosphate (ATP) |
| Cytosine (C) | Cytidine | Cytidine-5′-triphosphate (CTP) | |
| Guanine (G) | Guanosine | Guanosine-5′-triphosphate (GTP) | |
| Uracil (U) | Uridine | Uridine-5′-triphosphate (UTP) | |
| DNA | Adenine (A) | 2′-Deoxyadenosine | 2′-Deoxyadenosine-5′-triphosphate (dATP) |
| Cytosine (C) | 2′-Deoxycytidine | 2′-Deoxycytidine-5′-triphosphate (dCTP) | |
| Guanine (G) | 2′-Deoxyguanosine... |
Table of contents
- Cover image
- Title page
- Table of Contents
- Dedication
- Copyright
- Preface
- Chapter 1. RNA and the Cellular Biochemistry Revisited
- Chapter 2. Creating a Ribonuclease-Free Environment
- Chapter 3. RNA Isolation Strategies
- Chapter 4. The Truth About Tissues
- Chapter 5. Going Green: RNA and the Molecular Biology of Plants
- Chapter 6. Quality Control for RNA Preparations
- Chapter 7. cDNA—A Permanent Biochemical Record of the Cell
- Chapter 8. RT-PCR: A Science and an Art Form
- Chapter 9. Quantitative PCR Techniques
- Chapter 10. miRNA
- Chapter 11. RNA Interference and RNA Editing
- Chapter 12. Stringency: Conditions That Influence Nucleic Acid Structure
- Chapter 13. Electrophoresis of RNA
- Chapter 14. Photodocumentation and Image Analysis
- Chapter 15. Northern Analysis
- Chapter 16. Nucleic Acid Probe Technology
- Chapter 17. Isolation of Polyadenylated RNA
- Chapter 18. Quantification of Specific mRNAs by Nuclease Protection
- Chapter 19. Analysis of Nuclear RNA
- Chapter 20. Nonarray Methods for Global Analysis of Gene Expression
- Chapter 21. Genomes, Transcriptomes, Proteomes, and Bioinformatics
- Chapter 22. RNA Biomarker Discovery
- Chapter 23. High-Throughput Analysis of Gene Expression
- Chapter 24. Functional Genomics and Transcript Profiling
- Chapter 25. A Few RNA Success Stories
- Epilogue: A Few Pearls of Wisdom
- Appendix A. Maintaining Complete and Accurate Records
- Appendix B. Converting Mass to Moles
- Appendix C. Disposal of Ethidium Bromide and SYBR Green Solutions
- Appendix D. Removal of DNA From an RNA Sample
- Appendix E. Removal of RNA From a DNA Sample
- Appendix F. Useful Stock Solutions for the Molecular Biologist
- Appendix G. Silanizing Centrifuge Tubes and Glassware
- Appendix H. Dot Blot Analysis
- Appendix I. Electrophoresis: Principles, Parameters, and Safety
- Appendix J. Polyacrylamide Gel Electrophoresis
- Appendix K. Centrifugation as a Mainstream Tool for the Molecular Biologist
- Appendix L. Trypsinization Protocol for Anchorage-Dependent Cells
- Appendix M. Isolation of High-Molecular-Weight DNA by Salting-Out
- Appendix N. Phenol Preparation
- Appendix O. Deionization of Formamide, Formaldehyde, and Glyoxal
- Appendix P. Selected Suppliers of Equipment, Reagents, and Services
- Appendix Q. Useful SI Prefixes and Common SI Base Units
- Appendix R. Common Abbreviations
- Appendix S. Trademark Citations
- Glossary
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.4M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access RNA Methodologies by Robert E. Farrell Jr.,Robert E. Farrell, Jr. in PDF and/or ePUB format, as well as other popular books in Biological Sciences & Biology. We have over one million books available in our catalogue for you to explore.
