
- 552 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
About this book
This book provides an entry point into Systems Biology for researchers in genetics, molecular biology, cell biology, microbiology and biomedical science to understand the key concepts to expanding their work. Chapters organized around broader themes of Organelles and Organisms, Systems Properties of Biological Processes, Cellular Networks, and Systems Biology and Disease discuss the development of concepts, the current applications, and the future prospects. Emphasis is placed on concepts and insights into the multi-disciplinary nature of the field as well as the importance of systems biology in human biological research. Technology, being an extremely important aspect of scientific progress overall, and in the creation of new fields in particular, is discussed in 'boxes' within each chapter to relate to appropriate topics.
- 2013 Honorable Mention for Single Volume Reference in Science from the Association of American Publishers' PROSE Awards
- Emphasizes the interdisciplinary nature of systems biology with contributions from leaders in a variety of disciplines
- Includes the latest research developments in human and animal models to assist with translational research
- Presents biological and computational aspects of the science side-by-side to facilitate collaboration between computational and biological researchers
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Section II
Network Properties of Biological Systems
Chapter 3. Interactome Networks
Chapter 4. Gene Regulatory Networks
Chapter 5. Analyzing the Structure, Function and Information Flow in Signaling Networks using Quantitative Cellular Signatures
Chapter 6. Genetic Networks
Chapter 7. The Spatial Architecture of Chromosomes
Chapter 8. Chemogenomic Profiling
Chapter 9. Graph Theory Properties of Cellular Networks
Chapter 3
Interactome Networks
Chapter Outline
Introduction
Life Requires Systems
Cells as Interactome Networks
Interactome Networks and Genotype–Phenotype Relationships
Mapping and Modeling Interactome Networks
Towards a Reference Protein–Protein Interactome Map
Strategies for Large-Scale Protein–Protein Interactome Mapping
Large-Scale Binary Interactome Mapping
Large-Scale Co-Complex Interactome Mapping
Drawing Inferences from Interactome Networks
Refining and Extending Interactome Network Models
Predicting Gene Functions, Phenotypes and Disease Associations
Assigning Functions to Individual Interactions, Protein Complexes and Network Motifs
Towards Dynamic Interactomes
Towards Cell-Type and Condition-Specific Interactomes
Evolutionary Dynamics of Protein–Protein Interactome Networks
Concluding Remarks
Acknowledgements
References
Introduction
Life Requires Systems
What is Life? The answer to the question posed by Schrödinger in a short but incisive book published in 1944 remains elusive more than seven decades later. Perhaps a less ambitious, but more pragmatic question could be: what does Life require? Biologists agree on at least four fundamental requirements, among which three are palpable and easily demonstrable, and a fourth is more intangible (Figure 3.1). First, Life requires chemistry. Biomolecules, including metabolites, proteins and nucleic acids, mediate the most elementary functions of biology. Life also requires genes to encode and ‘reproduce’ biomolecules. For most organisms, cells provide the fundamental medium in which biological processes take place. The fourth requirement is evolution by natural selection. Classically, these ‘four great ideas of biology’ [1] have constituted the main intellectual framework around which biologists formulate biological questions, design experiments, interpret data, train younger generations of scientists and attempt to design new therapeutic strategies.
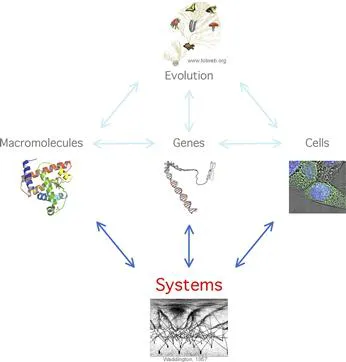
FIGURE 3.1 Systems as a fifth requirement for Life.
The next question then should be: even if we fully understood each of these four basic requirements of biology, would we be anywhere near a complete understanding of how Life works? Would we be able to fully explain genotype–phenotype relationships? Would we be able to fully predict biological behaviors? How close would we be to curing or alleviating suffering from human diseases? It is becoming clear that even if we knew everything there is to know about the four currently accepted requirements of biology, the answer to ‘What Life is’ would remain elusive.
The main reason is that biomolecules do not function in isolation, nor do cells, organs or organisms, or even ecosystems and sociological groups. Rather, biological entities are involved in intricate and dynamic interactions, thereby forming ‘complex systems’. In the last decade, novel biological questions and answers have surfaced, or resurfaced, pointing to systems as a fifth fundamental requirement for Life [2]. Although conceptual, systems may turn out to be as crucial to biology as chemistry, genes, cells or evolution (Figure 3.1).
Cells as Interactome Networks
The study of biological systems, or ‘systems biology’, originated more than half a century ago, when a few pioneers initially formulated a theoretical framework according to which multiscale dynamic complex systems formed by interacting biomolecules could underlie cellular behavior. To explain cellular differentiation, Delbrück hypothesized the existence of positive feedback circuits required for ‘bistability’, a model in which systems would remain stably activated after having been turned on, and conversely, remain steadily inactive once turned off [3]. Empirical evidence for feedback regulation in biology first emerged in the 1950s. The Umbarger and Pardee groups uncovered enzymatic feedback inhibition [4,5], and Novick and Weiner described the positive feedback circuit regulating the lac operon [6]. Monod and Jacob subsequently proposed how negative feedback circuits could account for homeostasis and other oscillatory phenomena observed in many biological processes [7]. These teleonomic arguments were later formalized by René Thomas and others in terms of requirements for cellular and whole organism differentiation based on positive and negative feedback circuits of regulation, using Boolean modeling as powerful simplifications of cellular systems [8] (see Chapter 10).
Equally enlightening theoretical systems properties were imagined beyond small-scale regulatory mechanisms composed of just a few molecules. Waddington introduced the metaphor of ‘epigenetic landscape’, whereby cells respond to genetic, developmental and environmental cues by following paths across a landscape containing peaks and valleys dictated by interacting genes and gene products [9]. This powerful idea, together with theoretical models of ‘randomly constructed genetic nets’ by Kauffman [10], suggested that a cellular system could be described in terms of ‘states’ resulting from particular combinations of genes, gene products, or metabolites, all considered either active or inactive at any given time. Complex wiring diagrams of functional and logical interconnectivity between biomolecules and genes acting upon each other could be imagined to depict how systems ‘travel’ from state to state over time throughout a ‘state space’ determined by intricate, sophisticated combinations of genotype, systems properties and environmental conditions. These concepts, elaborated at a time when the molecular components of biology were poorly described, remained largely ignored by molecular biologists until recently (see Chapter 15).
Over the past two decades, scientific knowledge of the biomolecular components of biology has dramatically increased. In particular, sequencing and bioinformatics have allowed prediction of coding and non-coding gene products at genome scale. Transcriptome sequencing approaches have revealed the existence of transcripts that had escaped prediction and which often remain of unknown function (see Chapter 2). Additionally, the list of known molecular components of cellular systems, including nucleic acids, gene products and metabolites, is lengthening and becoming increasingly detailed. With these advances came a humbling realization, best summed up as ‘too much data, too few drugs’ (see Chapter 8). It has become clearer than ever that knowing everything there is to know about each biomolecule in the cell is not sufficient to predict how the cell will react as a whole to particular external or internal perturbations.
Functional interactions, perhaps more so than individual components, mediate the fundamental requirements of the cell. Consequently, one needs to consider biological phenomena as the product of ensembles of interacting components with emergent properties that go beyond those of their individual components considered in isolation. One needs to step back and measure, model, and eventually perturb nearly all functional interactions between cellular components to fully understand how cellular systems work. In analogy to the word genome, the union of all interactions between all cellular components is termed the ‘interactome’. Our working hypothesis is that interactomes exhibit local and global properties that relate to biology in general, and to genotype–phenotype relationships in particular.
Interactome Networks and Genotype–Phenotype Relationships
Since drafts of a composite reference human genome sequence were released 10 years ago, powerful technological developments, such as next-generation sequencing, have started a true revolution in genomics [11–14]. With time, most human genotypic variations will be described, together with large numbers of phenotypic associations. Unfortunately, such knowledge cannot translate directly into new mechanistic understanding or therapeutic strategies, in part because the ‘one-gene/one-enzyme/one-function’ concept developed by Beadle and Tatum oversimplifies genotype–phenotype relationships [15]. In fact, Beadle and Tatum themselves state so in the introduction of their groundbreaking paper that initiated reductionism in molecular biology: ‘An organism consists essentially of an integrated system of chemical reactions controlled in some manner by genes. Since the components of such systems are likely to be interrelated in complex ways, it would appear that there must exist orders of directedness of gene control ranging from simple one-to-one relations to relations of great complexity.’
So-called complex traits provide the most compelling evidence of complexity between genotypes and phenotypes in human disease. Genome-wide association studies have revealed many more contrib...
Table of contents
- Cover image
- Title page
- Table of Contents
- Copyright
- List of Contributors
- Preface
- Reviewers
- Section I: Components of Biological Systems
- Section II: Network Properties of Biological Systems
- Section III: Dynamic and Logical Properties of Biological Systems
- Section IV: Systems and Biology
- Section V: Multi-Scale Biological Systems, Health and Ecology
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Handbook of Systems Biology by Marian Walhout,Marc Vidal,Job Dekker in PDF and/or ePUB format, as well as other popular books in Biological Sciences & Applied Mathematics. We have over 1.5 million books available in our catalogue for you to explore.