
eBook - ePub
Kingdoms and Domains
An Illustrated Guide to the Phyla of Life on Earth
- 732 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Kingdoms and Domains
An Illustrated Guide to the Phyla of Life on Earth
About this book
Now published by Academic Press and revised from the author's previous Five Kingdoms Third edition, this extraordinary, all inclusive catalogue of the world's living organisms describes the diversity of the major groups, or phyla, of nature's most inclusive taxa. Developed after consultation with specialists, this modern classification scheme is consistent both with the fossil record and with recent molecular, morphological and metabolic data. Generously illustrated, now in full color, Kingdoms and Domains is remarkably easy to read. It accesses the full range of life forms that still inhabit our planet and logically and explicitly classifies them according to their evolutionary relationships. Definitive characteristics of each phylum are professionally described in ways that, unlike most scientific literature, profoundly respect the needs of educators, students and nature lovers. This work is meant to be of interest to all evolutionists as well as to conservationists, ecologists, genomicists, geographers, microbiologists, museum curators, oceanographers, paleontologists and especially nature lovers whether artists, gardeners or environmental activists.Kingdoms and Domains is a unique and indispensable reference for anyone intrigued by a planetary phenomenon: the spectacular diversity of life, both microscopic and macroscopic, as we know it only on Earth today.
- New Foreword by Edward O. Wilson
- The latest concepts of molecular systematics, symbiogenesis, and the evolutionary importance of microbes
- Newly expanded chapter openings that define each kingdom and place its members in context in geological time and ecological space
- Definitions of terms in the glossary and throughout the book
- Ecostrips, illustrations that place organisms in their most likely environments such as deep sea vents, tropical forests, deserts or hot sulfur springs
- A new table that compares features of the most inclusive taxa
- Application of a logical, authoritative, inclusive and coherent overall classification scheme based on evolutionary principles
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Topic
Ciencias biológicasSubtopic
EcologíaSUPERKINGDOM EUKARYA
Origins by symbiogenesis
Organisms in this superkingdom composed of cells that contain more than two protein-coated 100 Å unit fibril Feulgen-stainable chromosomes per cell. The cells reproduce by mitosis. In mitosis pore-studded membrane-bounded nuclei totally or partially dissolve and re-form as two offspring nuclei. Display viable cytoplasmic and nuclear fusion and their reciprocal processes (for example, cytoplasmic and chromosomal doubling with subsequent meiotic or equivalent reduction of cytoplasmic and nuclear content; hence, Mendelian genetics). Cells have tubulin–actin cytoskeletons capable of intracellular motility visible in vivo. Cells contain kinetosome-centriole [9(3)+0] microtubular organizing centers or their derivatives. Organisms lack unidirectional gene (naked DNA) transfer except in the laboratory. There are at least two genomic ancestors to eukaryotic cells, one eubacterial and one archaebacterial.
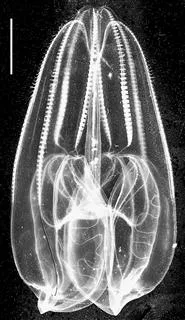
A-5A1 Bolinopsis infundibulum [Courtesy of M. S. Laverack.]
Superkingdom Eukarya
Kingdom Protoctista
Kingdom Animalia
Kingdom Fungi
Kingdom Plantae
Box Eukarya-i: Undulipodia from sulfur syntrophies
If the waste of one organism (sulfide, HS– in this case) becomes food or fuel of a second (a different kind of microbe that converts HS– to elemental sulfur, S) and then the sulfur can be used by the first organism to make more sulfide, we speak of a syntrophy (syn, together; trophy, feeding). Syntrophic partnerships are common in bacteria that tend to digest food outside of but near their bodies. The very first step in the origin of the first protoctists (and therefore the first nucleated cells) involved production of HS– by the archaebacterial ancestor, a wall-less, heat-tolerant cell much like today’s Thermoplasma acidophila (Searcy, 1980). We agree, but also posit that the partner in the syntrophy was an active swimming cell, a spirochete eubacterium that chemically converted low quantities of HS– to elemental sulfur. Because the swimming cell could tolerate only low concentrations of oxygen (<2%), the reaction HS–+oxygen=S for oxygen removal was crucial for its survival. Both partners sought organic compounds as food. Stuck together as a motility symbiosis, they tolerated ranges of oxygen from 0.000001% to about 5%. They maintained the association as they traversed regions of abundant sulfur compounds in search of food. Strong selective advantages to this sulfur syntrophy soon made it obligate; archaebacterial–eubacterial genomes merged, and membrane systems fused. The swimmer, we hypothesize, became the undulipodium (cilium or eukaryotic undulipodium with the well-studied [9(2+)+2] pattern of microtubules). From this versatile, wide-ranging partnership evolved the earliest intracellular motility, including the nucleus with mitotic cell division (“dance of the chromosomes”). The same organelle (the centriole or kinetosome) that gave rise to the microtubules of the undulipodium also became the spindle pole in mitosis; thus, the nucleus, centriole-kinetosome, and undulipodium formed the quintessentially eukaryotic organellar system called the karyomastigont. Origin of the karyomastigont was the crucial first step in the evolution of eukaryotes from prokaryotes. Other types of cell movement (phagocytosis, exocytosis, cyclosis, and so on), on this reckoning, evolved under very low atmospheric oxygen pressures before the incorporation of oxygen-respiring bacteria that became mitochondria and later, photosynthetic bacteria that became plastids.
The cells of animal, plant, fungal, and protoctist bodies, all composed of nucleated (“eukaryotic”) cells, tend to be larger and structurally more complex, sometimes vastly larger than those of any bacteria. However, the chemistry and fossil history of bacteria convince us that bacteria were the ultimate common ancestors of all extant life. Bacterial genes are made of DNA; bacterial cell membranes, invariably composed of fatty compounds linked to phosphoric acid and studded with proteins, are similar to those of eukaryotes.
What else took place during the evolutionary transition from bacterial cells to those of eukaryotes? Although some controversy persists over details, all scientists agree that nucleated cells (animal, plant, fungal, and protoctist) evolved not from any single hypertrophied ancestral bacterium, but from a mixed community of bacteria by way of mergers: literal incorporations of different kinds of bacteria. An α-proteobacterium that breathed oxygen became the mitochondrion, and photosynthetic cyanobacteria became the chloroplasts and all other plastids. At least two very different bacterial lineages had already fused in the formation of ancestors of all eukaryotes. We suspect that these were archaebacteria, which converted sulfur to sulfide, and eubacteria, common spirochetes that converted sulfide to sulfur for oxygen protection. Animal, plant, and other eukaryotic cells retain sulfur metabolism similar to today’s archaebacteria. The cytoplasm of all eukaryotes investigated still produces HS–. Hydrogen sulfide peaks just before mitosis and declines afterward as measured in some ciliates and even reversibly puts mice to sleep when given in tiny amounts. Our ancient legacy of sulfur syntrophy is still with us today.
References
Searcy, D. G., and R. J. Delange, Thermoplasma acidophilum histone like protein: Partial amino acid sequence suggestive of homology to eukaryotic histones. Biochim. Biophys. Acta 609:197–200; 1980a.
Searcy, D. G., and D. B. Stein, Nucleoprotein subunit structure in an unusual prokaryotic organism: Thermoplasma acidophilum. Biochim. Biophys. Acta 609:180–195; 1980b.
Box Eukarya-ii: Eukaryosis—Life histories, life cycles
The three major kingdoms of eukaryotes—fungi, plants, and animals—can be distinguished by the position of meiosis in their life cycles: fungi are haploid with zygotic meiosis, plants alternate haploidy (one set of chromosomes) and diploidy (two sets of chromosomes) with sporogenic meiosis, and animal bodies are diploid with somatic meiosis. The “imperfections and oddities” Darwin cited as illustrative of “descent with modification” are in the protoctists, which show every sort of variation on life cycle pattern from no meiosis-fertilization cycle at all to the complexity of the foraminifera with every sort of possibility. We speak of life cycle when we know ploidy of the cells, but only of life history when our information is limited to morphological data without karyology (Figures Eukarya-ii-1 through Eukarya-ii-41,2,3,4Figures Eukarya-ii-1 through Eukarya-ii-41,2,3,4)
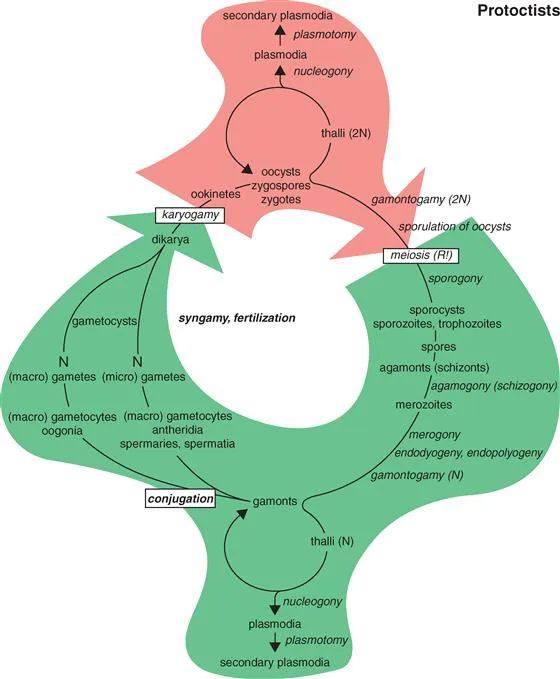
Figure Eukarya-ii-1 Generalized protoctist life cycle. Meiosis gives rise to haploid nuclei in cells of organisms. These occur e.g., in Apicomplexa (Pr-7) as resistant sporocysts, motile sporozoites or feeding trophozoites. Depending on environmental conditions a haploid cell or multicellular organism may remain in a uniparental, trophic or reproductive state as a haploid agamont (if it reproduces before it makes gametes). Or by mitotic growth and differentiation it may become a gamont. A gamont is an organism, either haploid or diploid, that by mitosis or meiosis respectively, makes gamete nuclei or gamete cells. The haploid organism may differentiate reproductive thalli, plasmodia, pseudoplasmodia or other structures without meiosis and remain an agamont. The haploid may form egg-producing oogonia, sperm-filled antheridia or develop isogametous (look-the-same) gametes in which case it changes, by definition, from an agamont to a gamont. Protoctist generative nuclei or cells may also remain in the diploid state and grow large and/or reproduce by multiple fission, hyphae, plasmodia, thalli, spores or other agamontic life history forms. Some diploid nuclei undergo meiosis in uni- or multicellular protoctists to produce more offspring as agamonts, gamonts or gametes. Gametes may be haploid nuclei only (as in some ciliates and foraminifera) or whole gamont bodies (as in many sexual algae or water molds. Gamontogamy, cytogamy and/or karyogamy (= conjugation, sexual fusion of cytoplasm of gaemete-formers or their gam...
Table of contents
- Cover Image
- Content
- Title
- Copyright Page
- Dedication
- List of Figures
- List of Tables
- Foreword
- Foreword To 1st–3rd editions
- Preface
- Acknowledgments
- Introduction
- SUPERKINGDOM PROKARYA
- SUPERKINGDOM EUKARYA
- General glossary
- Organism Glossary
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Kingdoms and Domains by Lynn Margulis,Michael J. Chapman,Michael J Chapman in PDF and/or ePUB format, as well as other popular books in Ciencias biológicas & Ecología. We have over 1.5 million books available in our catalogue for you to explore.