
eBook - ePub
Biodiversity of Fungi
Inventory and Monitoring Methods
- 777 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
About this book
Biodiversity of Fungi is essential for anyone collecting and/or monitoring any fungi. Fascinating and beautiful, fungi are vital components of nearly all ecosystems and impact human health and our economy in a myriad of ways. Standardized methods for documenting diversity and distribution have been lacking. A wealth of information, especially regrading sampling protocols, compiled by an international team of fungal biologists, make Biodiversity of Fungi an incredible and fundamental resource for the study of organismal biodiversity.
Chapters cover everything from what is a fungus, to maintaining and organizing a permanent study collection with associated databases; from protocols for sampling slime molds to insect associated fungi; from fungi growing on and in animals and plants to mushrooms and truffles. The chapters are arranged both ecologically and by sampling method rather than by taxonomic group for ease of use. The information presented here is intended for everyone interested in fungi, anyone who needs tools to study them in nature including naturalists, land managers, ecologists, mycologists, and even citizen scientists and sophiscated amateurs.
- Covers all groups of fungi - from molds to mushrooms, even slime molds
- Describes sampling protocols for many groups of fungi
- Arranged by sampling method and ecology to coincide with users needs
- Beautifully illustrated to document the range of fungi treated and techniques discussed
- Natural history data are provided for each group of fungi to enable users to modify suggested protocols to meet their needs
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Topic
Ciencias biológicasSubtopic
BiologíaPART I
GENERAL ISSUES
1 FUNGI AND THEIR ALLIES
A BROAD VIEW OF EUKARYOTES
KINGDOM FUNGI
PHYLUM CHYTRIDIOMYCOTA (ZOOSPORIC FUNGI)
PHYLUM ZYGOMYCOTA
CLADE GLOMALES
PHYLUM ASCOMYCOTA
Class Archiascomycetes
Class Saccharomycetes
Class Euascomycetes
PHYLUM BASIDIOMYCOTA
Class Ustilaginiomycetes
Class Urediniomycetes
Class Hymenomycetes
KINGDOM STRAMINIPILA (HETEROKONT ZOOSPORIC ORGANISMS)
Oomycota
Hyphochytriomycetes
Labyrinthulales and Thraustochytriales (and Aplanochytrium)
SLIME MOLDS
Plasmodiophorales (Parasitic Slime Molds)
Myxomycetes, Protostelids, and Dictyostelids (Plasmodial and Cellular Slime Molds)
Acrasid Slime Molds
CONCLUSIONS
Fungi are heterotrophic organisms that permeate our environment. With few exceptions fungi have filamentous bodies enclosed by cell walls, are nonmotile, and reproduce both sexually and asexually by spores. During the last decade, mycologists have made unprecedented progress toward producing a phylogenetic classification of fungi; a skeleton phylogeny based on analyses of DNA characters was developed relatively early on (Bruns et al. 1991; Alexopoulos et al. 1996; McLaughlin et al. 2001). Such a phylogeny serves as a basis on which hypotheses of fungal evolution can be developed. One important finding has been that “fungi” are polyphyletic (Table 1.1); their morphologies are convergent, having been derived independently from among several independent eukaryotic lineages (Fig. 1.1). A monophyletic group, exclusive of slime molds and oomycetes, is well defined and supported as “true fungi,” a kingdom-level taxon (Barr 1992; Bruns et al. 1993; Baldauf et al. 2000; Keeling et al. 2000).
TABLE 1.1 Fungi and Fungus-like Organisms Previously Classified as Fungi*
| Fungi |
| Chytridiomycota |
| Zygomycota |
| Ascomycota |
| Basidiomycota |
| Straminipila |
| Oomycota |
| Hyphochytriomycota |
| Labyrinthulales |
| Thraustochytriales |
| Slime molds |
| Plasmodiophorales |
| Myxomycota |
| Dictyosteliomycota |
| Acrasiomycota |
* Taxonomic ranks of the clades are not indicated (see text).
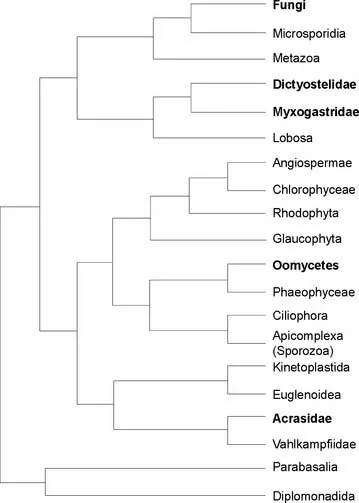
FIGURE 1.1 Phylogenetic positions of organisms studied by mycologists (bold text). The tree is unrooted, and all organisms are eukaryotic (i.e., they possess nuclei and other membrane-bounded organelles). The tree is based on phylogenetic analyses of sequences of up to four protein-coding (α-tubulin, β-tubulin, actin, and elongation factor 1-alpha [EF-1α]) regions in addition to rDNA.
(Based on Baldauf et al. 2000.)
Historically fungi have been compared with plants and included in the study of botany. Contemporary studies, however, indicate that members of Kingdom Fungi are most closely related to animals, not plants, possibly through a choanoflagellate-like ancestor (Barr 1992; Baldauf and Palmer 1993; Wainright et al. 1993; Ragan et al. 1996; Baldauf et al. 2000; Cavalier-Smith 2001). The phylogenetic positions of other groups once considered as fungi are not always well defined among eukaryotes (see “Kingdom Straminipila” and “Slime Molds,” later in this chapter).
Thus far, fewer than 800 fungi or fungus-like taxa have been included together in any single phylogenetic reconstruction (Tehler et al. 2000), a number representing less than 1% of the 80,000 currently listed species (see Hawksworth et al. 1995). Estimates of up to 1.5 million species of fungi (six times more than the number of recognized species of land plants) are daunting; the missing fungi must be discovered, identified, and incorporated into the taxonomic framework. In addition to the relatively low numbers of taxa included in the trees, taxonomic coverage across higher taxa is limited. Until now only molecular characters from ribosomal RNA genes (rDNA) have been used widely in phylogenetic studies of fungi. The recent increase in use of other DNA regions and incorporation of phenotypic characters to resolve conflicting trees based on single genes and analytical approaches is encouraging.
Fungal classification, as opposed to phylogeny, is in a state of flux. Ideally, classifications should reflect phylogenetic relationships, but the relationships of all the groups covered in this volume have not been resolved nor have all of the groups been represented in analyses. Thus, the formal and complex classification systems that have been devised (e.g., Cavalier-Smith 2001) may vary in their inclusiveness. Systematists who study many types of organisms face this situation. Nevertheless, one reactionary suggestion called for a moratorium on publication of higher-level taxonomic systems until 2050 (Karatygin 1999). Hibbett and Donoghue (1998) suggested a more practical solution—the use of clade names rather than formal taxon designations. We use this approach, especially for higher taxa, in the following discussion.
Nowhere are examples of convergent evolution more evident than among organisms occupying the same habitat, which are grouped together for treatment in this volume. In this chapter we review the current state of knowledge of the phylogeny of true fungi (Kingdom Fungi) and fungus-like organisms placed in several additional distinct clades (Table 1.1). The two-volume work edited by McLaughlin et al. (2001) provides detailed information on fungi and fungus-like organisms; only a few of the chapters are cited here.
A BROAD VIEW OF EUKARYOTES
Previous studies based on small subunit rRNA genes (SSU rDNA) suggest that it may not be possible to reconstruct patterns of an ancient rapid radiation of the so-called “crown eukaryotes” (see “Kingdom Fungi,” later in this chapter), especially when based solely on characters from rDNA. Baldauf and colleagues (Baldauf et al. 2000), in contrast, used multiple genes and partitioning of data to develop a hypothesis of the relationships of the large diverse group of eukaryotes, which includes fungi and fungus-like organisms (Fig. 1.1). Their study differed from others in its approach because Baldauf and colleagues (2000) analyzed deduced amino acid sequences from four protein-coding regions (α-tubulin, β-tubulin, actin, and elongation factor 1-alpha [EF-1α]) in addition to rDNA. Consequently, they obtained better resolution of the deep branches in the tree (that is, those that represent the earliest divergences over a great period).
Many of the relationships proposed in Baldauf and colleagues’ (2000) study were not entirely new; however, the large data set they used supported several controversial or conflicting tree branches, depending on their use of either rDNA or protein-coding genes. Relationships identified or strengthened in their phylogenetic analyses included the Fungi-Microsporidia link, the grouping of dictyostelid and plasmodial slime molds, and the remote basal position of the acrasid slime molds. In some instances, correspondence of non-molecular traits (e.g., flagellum type) and arrangement of mitochondrial cristae provided additional support. Taking a broad view of fungi and fungus-like organisms and using a wide variety of characters, Baldauf and colleagues (2000) derived the following relationships (Fig. 1.1):
• Kingdom Fungi, (traditionally the phyla Ascomycota, Basidiomycota, Zygomycota, and Chytridiomycota; see “Kingdom Fungi,” later in this chapter) comprises a monophyletic group of “true” Fungi.
• Microsporidia are a sister group to Fungi, and these two taxa form a monophyletic grouping with the Metazoa (animals) linked by a choanoflagellate-like common ancestor.
• Although still scantily sampled, dictyostelid...
Table of contents
- Cover image
- Title page
- Table of Contents
- Copyright
- FOREWORD
- PREFACE
- ADDRESSES OF AUTHORS AND CONTRIBUTORS
- INTRODUCTION
- PART I: GENERAL ISSUES
- PART IIA: RECOMMENDED PROTOCOLS FOR SAMPLING PARTICULAR GROUPS OF FUNGI: DIRECT COLLECTING AND ISOLATION PROTOCOLS FOR MACROFUNGI AND MICROFUNGI ON SOIL, WOOD, LEAVES, LICHENS, AND OTHER SUBSTRARA
- PART IIB: RECOMMENDED PROTOCOLS FOR SAMPLING PARTICULAR GROUPS OF FUNGI: ISOLATION PROTOCOLS FOR READILY CULTURABLE MICROFUNGI ASSOCIATED WITH PLANTS AND FUNGI
- PART IIC: RECOMMENDED PROTOCOLS FOR SAMPLING PARTICULAR GROUPS OF FUNGI: COLLECTING AND ISOLATION PROTOCOLS FOR FUNGI ASSOCIATED WITH ANIMALS
- PART IID: RECOMMENDED PROTOCOLS FOR SAMPLING PARTICULAR GROUPS OF FUNGI: COLLECTING AND ISOLATION PROTOCOLS FOR AQUATIC FUNGI AND FOR PROTOCTISTANS FORMERLY TREATED AS FUNGI
- PART III: APPENDICES, GLOSSARY, LITERATURE CITED, AND INDEX
- GLOSSARY
- LITERATURE CITED
- INDEX
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Biodiversity of Fungi by Mercedes S. Foster,Gerald F. Bills in PDF and/or ePUB format, as well as other popular books in Ciencias biológicas & Biología. We have over 1.5 million books available in our catalogue for you to explore.