
- 414 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Cellular Endocrinology in Health and Disease
About this book
Cellular Endocrinology in Health and Disease describes the underlying basis of endocrine function, providing an important tool to understand the fundamentals of endocrine diseases. Delivering a comprehensive review of the basic science of endocrinology, from cell biology to human disease, this work explores and dissects the function of a number of cellular systems. Among these are those whose function was not obvious until recently, including the endocrine functions of bone and the adipose tissue.
Providing content that crosses disciplines, Cellular Endocrinology in Health and Disease details how cellular endocrine function contributes to system physiology and mediates endocrine disorders. A methods section proves novel and useful approaches across research focus that will be attractive to medical students, residents, and specialists in the field of endocrinology, as well as to those interested in cellular regulation. Editors Alfredo Ulloa-Aguirre and P. Michael Conn, experts in molecular and cellular aspects of endocrinology, deliver contributions carefully selected for relevance, impact, and clarity of expression from leading field experts.
- Covers systemic endocrine action at the cellular level in both health and disease
- Delivers information on the integration of cell identity and endocrinology
- Incorporates recent developments in endocrinology to provide an up-to-date reference to researchers
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Topic
MedicinaSubtopic
Endocrinologia e metabolismoChapter 1
Thyroid Hormone Receptors and their Role in Cell Proliferation and Cancer
Olaia Martínez-Iglesias, Lidia Ruiz-Llorente, Constanza Contreras Jurado and Ana Aranda, Instituto de Investigaciones Biomédicas “Alberto Sols,” Madrid, Spain, Consejo Superior de Investigaciones Científicas and Universidad Autónoma de Madrid, Madrid, Spain
The thyroid hormone receptors, TRα and TRβ, are ligand-dependent transcription factors that regulate gene expression by recruitment of coactivators and corepressors. These receptors play an important role in normal and malignant cell proliferation. Particularly, we have shown that TRs antagonize ras-dependent proliferation, transformation and tumorigenesis in fibroblasts. Furthermore, expression of TRβ in human cancer cells retards tumor growth and inhibits invasiveness, extravasation and metastasis formation in euthyroid nude mice. When cells are inoculated into hypothyroid hosts, tumor growth is retarded, but tumors that do grow are more invasive and metastatic growth is enhanced. Increased malignancy of skin tumors is found in mice lacking TRs, further demonstrating the role of these receptors as inhibitors of tumor progression and suggesting that they represent a potential therapeutic target in cancer. TRs have a dual effect on proliferation, because they are required for proliferation of normal hepatocytes or keratinocytes, while acting as inhibitors of tumor progression.
Keywords
Coactivators; Corepressors; Metastasis; Proliferation; Thyroid hormone receptors; Tumor progression
Thyroid Hormone Action
The important physiological actions of the thyroid hormones (THs) are mediated by binding to the nuclear thyroid hormone receptors (TRs). The thyroid gland produces predominantly thyroxine (T4), but triiodothyronine (T3) is the most active TH, since it has a higher affinity by the receptors.1 THs are released by the thyroid gland to the bloodstream and they enter the cells through the adenosine triphosphate (ATP)-dependent monocarboxylate transporters MCT8 and MCT10 and the organic anion transporter proteins (OATPs).2 The amount of T3 available for binding to the nuclear receptors is regulated by cell-specific expression of selenoenzymes deiodinases (DIOs). DIO1 and DIO2 catalyze the conversion of T4 to T3 in target tissues, increasing intracellular levels of the active hormone, while DIO3 causes hormone inactivation since it converts T4 and T3 by inner ring deiodination to the inactive metabolites reverse T3 (rT3) and T2, respectively.
TRs belong to the superfamily of nuclear receptors and act as ligand-dependent transcription factors.3 Several TR protein isoforms are generated by promoter use or alternative splicing of the primary transcripts of the TRα and TRβ genes. The TRα1, TRβ1 and TRβ2 are the main hormone-binding isoforms and their relative levels of expression vary among cell types and at different developmental stages, suggesting that they could have organ-specific functions. In the case of TRβ, TRβ1 is more widely expressed, while the expression of TRβ2 is restricted to the anterior pituitary, and some neural cells.4,5 Studies with genetically modified mice have shown that TRα and TRβ can substitute for each other to mediate some actions of the thyroid hormones but they can also mediate isoform-specific functions.6
As shown in Figure 1.1, TRs are composed of several functional domains. The N-terminal region (A/B) contains a constitutive ligand-independent transcriptional activation domain, the autonomous activation function 1 (AF-1). This region is followed by the DNA-binding domain (DBD), or region C. The DBD is the most conserved region among the nuclear receptors and is composed of two zinc fingers. In each zinc finger, four invariable cysteines coordinate tetrahedrically with one zinc ion. Amino acids required for discrimination of the thyroid hormone response element (TRE) are present at the base of the first finger in a region termed the “P box,” and other residues of the second zinc finger that form the so called “D box” are involved in dimerization. Through the DBD the receptors interact with the major groove of DNA. A hinge domain, or D region, connects the DBD with the E region or ligand-binding domain (LBD), also responsible for dimerization. This hinge domain contains residues essential for interaction with corepressors. Crystallographic analysis has shown that the LBDs are formed by 12 α-helices, and the C-terminal helix (H12) encompasses the ligand-dependent transcriptional activation function, or AF-2.
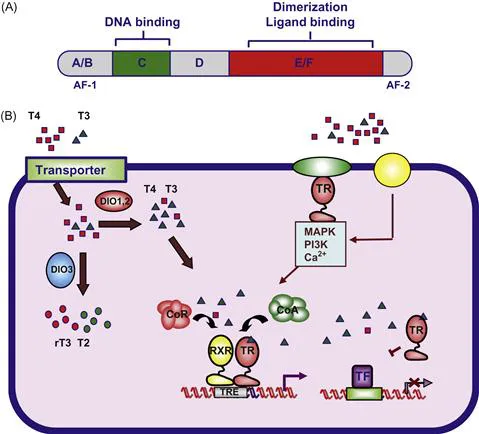
Figure 1.1 Mechanism of action of the thyroid hormone receptors.
(A) Schematic representation of a thyroid hormone receptor, showing the different functional domains. (B) Thyroxine (T4) and triiodothyronine (T3) enter the cell through transporter proteins such as MCT8 and 10 or OATPs. Inside the cells, deiodinases (DIO1,2) convert T4, to the more active form T3. DIO3 produces rT3 and T2 from T4 and T3, respectively. T3 binds to nuclear thyroid hormone receptors (TRs) that regulate transcription by binding, generally as heterodimers with the retinoid X receptor (RXR), to positive or negative thyroid hormone response elements (TREs) located in regulatory regions of target genes. Activity is regulated by an exchange of corepressor (CoR) and coactivator (CoA) complexes. TRs can also regulate the activity of genes that do not contain a TRE through “cross-talk” with other transcription factors (TF) that stimulate target gene expression. Binding of T3 to a subpopulation of receptors located outside the nuclei can also cause rapid “non-genomic” effects through interaction with adaptor proteins, leading to stimulation of signaling pathways. T4 can also bind to putative membrane receptors such as integrin αVβ3 inducing mitogen activated protein kinase (MAPK) activity.
(A) Schematic representation of a thyroid hormone receptor, showing the different functional domains. (B) Thyroxine (T4) and triiodothyronine (T3) enter the cell through transporter proteins such as MCT8 and 10 or OATPs. Inside the cells, deiodinases (DIO1,2) convert T4, to the more active form T3. DIO3 produces rT3 and T2 from T4 and T3, respectively. T3 binds to nuclear thyroid hormone receptors (TRs) that regulate transcription by binding, generally as heterodimers with the retinoid X receptor (RXR), to positive or negative thyroid hormone response elements (TREs) located in regulatory regions of target genes. Activity is regulated by an exchange of corepressor (CoR) and coactivator (CoA) complexes. TRs can also regulate the activity of genes that do not contain a TRE through “cross-talk” with other transcription factors (TF) that stimulate target gene expression. Binding of T3 to a subpopulation of receptors located outside the nuclei can also cause rapid “non-genomic” effects through interaction with adaptor proteins, leading to stimulation of signaling pathways. T4 can also bind to putative membrane receptors such as integrin αVβ3 inducing mitogen activated protein kinase (MAPK) activity.
TRs regulate gene transcription by binding, preferentially as heterodimers with retinoid X receptors (RXRs), to short DNA binding motifs, called thyroid hormone response elements or TREs, which are located in regulatory regions of target genes.7 TREs are composed of two copies of the AGG/TTCA motif. They can be configured as palindromes (Pal), inverted palindromes (IPs), or direct repeats spaced preferably by four non-conserved nucleotides (DR4). Although TRs can bind to their response elements as monomers or homodimers, heterodimerization with RXR strongly increases the affinity for DNA and transcriptional activity.
Transcriptional regulation by these receptors is mediated by the recruitment of coactivators and corepressors.3,8,9 In the absence of ligand, TRs can act as constitutive repressors when bound to TREs, due to their association with corepressors such as NCoR (nuclear receptor corepressor) or SMRT (silencing mediator of retinoic and thyroid receptor). NCoR and SMRT belong to multicomponent repressor complexes that contain histone deacetylases (HDACs) and cause chromatin compaction and consequently transcriptional inhibition.10 NCoR and SMRT are related both structurally and functionally. They contain three autonomous repressor domains (RD) and a receptor interacting domain (the CoRNR motif) located toward the carboxyl terminus. Transcriptional repression by the corepressor-bound receptors appears to be mediated by the recruitment of HDACs to the target gene. HDAC1 or 2 (class I deacetylases) are recruited to the first RD of the corepressors via the adaptor mSin3 protein, and the RD3 has been demonstrated to repress transcription by directly interacting with class II deacetylases (HDACs 4, 5 and 7). In addition, a repressor complex containing the corepressors, HDAC3 and transducin beta-like proteins (TBL1 or TBL1R) appears to be required for repression by TR. Although a receptor CoR box, located within the hinge region, is essential for interaction of receptors with the corepressors, the CoRNR motif does not interact directly with residues in this region, but docks to a hydrophobic groove in the surface of the LBD at H3 and 4.
Hormone binding induces a conformational change in the receptor that allows the release of corepressors and allows the recruitment in a sequential manner of coactivator complexes. The stronger change observed in the receptors upon ligand binding is the position of H12.11 This helix projects away from the body of the LBD in the absence of ligand. However, upon hormone binding H12 moves in a “mouse-trap” model being tightly packed against H3 or 4 and making direct contacts with the ligand. This change generates a hydrophobic cleft responsible for interaction with coactivators.12 A glutamic acid residue in H12 and a lysine residue in H3, which are conserved throughout the superfamily of nuclear receptors, interact directly with the coactivator and form a charge clamp that stabilizes binding. Consequently, mutation of these residues abolishes coactivator binding and causes the loss of thyroid hormone-dependent transcriptional activation.13 Since the coactivator binding surface overlaps with that involved in corepressors interaction, coactivator and corepressor binding is mutually exclusive. Some coactivators belong to ATP-dependent chromatin-remodeling complexes, others are part of complexes that induce post-translational modifications of histones, such as acetylation or arginine methylation, and others interact with the basic transcriptional machinery causing the recruitment of RNA polymerase II to the target promoter. Binding of the coactivators causes chromatin decompaction and transcriptional activation.
In addition to causing ligand-dependent transcriptional activation, TRs can also repress gene transcription in a hormone-dependent manner. In some cases, this repression is associated with binding to negative TREs (nTREs). Although the properties of nTREs are not yet well known, these elements are often located very close to the transcriptional start site,3 and corepressors and deacetylase activity appear to be involved in hormone-dependent negative regulation.14 TRs can also regulate the expression of genes that do not contain a TRE by positive or negative interference with the activity of other transcription factors or signaling pathways, a mechanism referred to as transcriptional crosstalk.3 Thus, we have shown that TRs can antagonize AP-1,15,16 cyclic AMP (cAMP) response element-binding protein (CREB),17,18 or NF-kB-mediated transcription.19,20 In this case, the receptors do not bind directly to the DNA recognition elements for these transcription factors in the target gene, but can be tethered to these binding motifs via protein-to-protein interactions. This type of transcriptional crosstalk between transcription factors and nuclear receptors has been shown to be critical for regulation of many cellular functions, including anti-inflammatory and anti-proliferative actions of nuclear receptor ligands.21–23 Finally, thyroid hormones can elicit rapid non-genomic effects initiated at the cell membrane that can lead to stimulation of kinase pathways. These actions could be mediated by a fraction of membrane-associated nuclear receptors, or by occupancy of putative membrane receptors, such as integrin αVβ3, which would bind T4 preferentially.9 Figure 1.1B illustrates the main aspects of thyroid hormone actions on cells.
TRs and Cancer
The first evidence linking TRs with cancer was the finding that TRα is the cellular counterpart of the v-erbA oncogene of the avian erythroblastosis virus (AEV), a retrovirus that causes erythroleukemia and sarcoma in chickens. v-ErbA acts as a constitutive dominant-negative of TRs since it contains mutations that abolish ligand binding, recruitment of coactivators and hormone-dependent transcriptional stimulation, while maintaining the ability to bi...
Table of contents
- Cover image
- Title page
- Table of Contents
- Copyright
- List of Contributors
- Preface
- Chapter 1. Thyroid Hormone Receptors and their Role in Cell Proliferation and Cancer
- Chapter 2. The Molecular Cell Biology of Anterior Pituitary Cells
- Chapter 3. Sensing Calcium Levels: The Biology of the Parathyroid Cell
- Chapter 4. The Biology of Pituitary Stem Cells
- Chapter 5. The Na+/I− Symporter (NIS) and Thyroid Hormone Biosynthesis
- Chapter 6. The Follicle-Stimulating Hormone Signaling Network in Sertoli Cells
- Chapter 7. Epigenetics of Pituitary Cell Growth and Survival
- Chapter 8. Epigenetic and Developmental Basis of Risk of Obesity and Metabolic Disease
- Chapter 9. Unraveling the Mechanism of Action of the GnRH Pulse Generator: A Possible Role for Kisspeptin/Neurokinin B/Dynorphin (KNDy) Neurons
- Chapter 10. Proteomics in Reproduction: The Dialogue Between the Blastocyst and the Endometrium
- Chapter 11. Transcriptome Analysis of Adrenocortical Cells in Health and Disease
- Chapter 12. Bone as an Endocrine Organ
- Chapter 13. Regulation of Steroidogenesis
- Chapter 14. Adipose Tissue as an Endocrine Organ
- Chapter 15. Insulin-Secreting Cell Lines: Potential for Research and Diabetes Therapy
- Chapter 16. Architecture and Morphology of Human Pancreatic Islets
- Chapter 17. Computational Models to Decipher Cell-Signaling Pathways
- Chapter 18. Defects in Ovarian Steroid Hormone Biosynthesis
- Chapter 19. Control of the GnRH Pulse Generator
- Chapter 20. Endocrinology of the Single Cell: Tools and Insights
- Chapter 21. Intracellular Trafficking of G Protein-Coupled Receptors to the Plasma Membrane in Health and Disease
- Chapter 22. Iodothyronine Deiodinases: Emerging Clinical Crossroads
- Chapter 23. MicroRNAs and Long Non-Coding RNAs in Pancreatic Beta Cell Function
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Cellular Endocrinology in Health and Disease by Alfredo Ulloa-Aguirre,P. Michael Conn in PDF and/or ePUB format, as well as other popular books in Medicina & Endocrinologia e metabolismo. We have over 1.5 million books available in our catalogue for you to explore.