The potential impact of anthropogenic pollutants such as agrochemicals on the environment is of global concern. Increasing use of certain compounds can result in contamination of food, water and atmospheric systems and in order to combat this pollution it is important to be able to accurately monitor the short and long term effects.
This book describes the latest aquatic species models used as indicators of the toxic effects of environmental pollutants, including models that have not routinely been used. The book enables understanding of the effects of pollutants in non-target species, and therefore enables analysis of the effects on ecosystems.
This book will be of interest to anyone interested in developing new biomarker species with high degrees of ecological relevance. It will serve as a useful resource for regulatory and research toxicologists, particularly those studying freshwater, marine water and sediment environments.

- 538 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
About this book
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Topic
Physical SciencesSubtopic
EcologyCHAPTER 11
The Use of Fish as Model Aquatic Organisms in Genotoxicity Studies
Uludag University, Faculty of Arts & Sciences Department of Biology, 16059, Bursa, Turkey
E-mail: [email protected]
11.1 Introduction
As a result of the increasing input of genotoxic agents into environment, growing attention has recently been paid to the assessment of the effect of genotoxic different agents on organisms and ecosystems. Agents that induce alterations in genetic material are called as genotoxic agents or genotoxins. Genetic toxicology studies have mainly two aims: (1) to detect and analyze the potential harmful effects of physical and chemical agents capable of damaging DNA; and (2) to investigate the mechanisms of action of those agents. Exposure to genotoxic agents might result in changes in DNA sequences and, if not repaired quickly and correctly, these changes can lead to heritable changes i.e. mutations, which may lead to cancer. Mutations may affect the organism itself through changes in body cells, or they may be passed on to other generations through alteration of the germ cells (Figure 11.1).
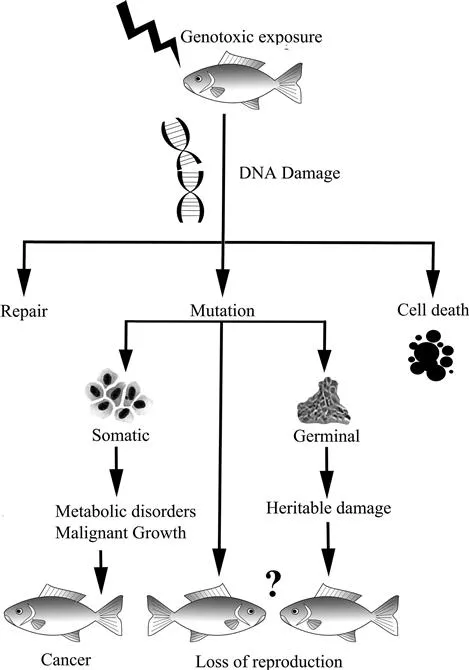
Figure 11.1 Schematic presentation of anticipated consequences of genotoxic exposure in fish.
The aquatic environment is the ultimate recipient of man-made contaminants, i.e. industrial, agricultural and domestic wastes. At this point, fish serve as the best model for detecting genotoxic effects in the aquatic environment. The assessment of the ecological risks associated with environmental genotoxic exposure is usually based on individual responses. Genotoxic damage induced in somatic cells can lead to cell death or malignancies, which might be manifested as different types of disorders in the organism itself. On the other hand, some genotoxins are prone to affect offspring generations directly or indirectly through the germ cells and reproductive process, which may eventually cause loss of reproduction and changes in population dynamics.1,2
11.2 Fish as Model Organisms in Genetic Toxicology
Fish may act as sentinel organisms in evaluation of genotoxicity as they often respond to toxicants in a manner similar to higher vertebrates.3,4 They are capable of inhabiting different environmental zones in aquatic habitats. Teleost fish, especially small aquarium species, can be easily held in the laboratory and exposed to chemicals under controlled conditions. Furthermore, they therefore offer a large number of models of adaptation and response to a wide variety of natural and anthropogenic environmental conditions. Another advantage of using fish as a model organism is that they are directly and effectively exposed to waterborne contaminants through their gills and gastro-intestinal system. Fish can be used to screen for chemicals that have the potential to induce genotoxic damage or carcinogenic effects in humans and other organisms or to determine the distribution and effects of pollutants in the aquatic environment. Thus, especially within the last two decades, the use of fish as suitable models for genotoxicity monitoring in aquatic environments has become popular. Literature surveys indicate that more than 40 marine and freshwater fish species have been used to assess genotoxicity, including widespread, endemic, tropical or even aquarium fish, mainly depending on the location and availability of specimens. Some of the most commonly used model fish species are listed in Table 11.1.
Table 11.1 The most commonly used model fish species in aquatic genotoxicity studies. Images used from references.5–14

In aquatic genotoxicity studies, the main cell types used in fish are: peripheral erythrocytes, gill epithelial cells, liver hepatocytes, kidney cells and fin epithelial cells (Figure 11.2). Among these cells, the use of erythrocytes has gained importance in biomonitoring of aquatic genotoxicity in fish thanks to their nucleated structure as well as the practicality of obtaining and processing blood samples. The use of fin and blood cells has further advantages over other cell types as it is not necessary to kill fish to obtain these cell samples.
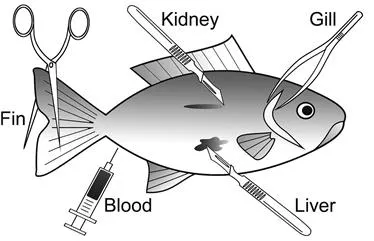
Figure 11.2 The main fish tissues used in genotoxicity assessments (drawing by T. Cavas).
11.3 Genotoxicity Techniques
The genotoxicity of different factors in fish has been assessed by using different test systems. These assays differ in their end points and specificities to explore the genotoxic effects induced by toxicants. In this chapter, some of the most commonly used in vivo genotoxicity techniques are described, namely chromosome aberrations, micronucleus, nuclear abnormalities and the Comet assay.
11.3.1 Cytogenetic Techniques
Genotoxicity studies in fish started with the use cytogenetic techniques originally developed for mammals.15–18 However, the use of metaphase techniques, such as sister chromatid exchange (SCE) and chromosomal aberrations (CA), is not practical for most fish species and only a limited number of fish species have suitable chromosomes for metaphase analyses. This is mainly because the karyotype of most fish consists of a large number of small and irregular chromosomes. Furthermore, karyological data is available for about only 10% of approximately 25 000 taxonomically known species and individual specimens with different karyotype compositions can be seen in the same species or population.19–21 Therefore, a fish species to be used should have a well-established karyotype with a small number of large chromosomes. Some fish species with a suitable karyotype that can be used in chromosomal aberration analyses are given in Table 11.2. On the other hand, the micronucleus test provides an alternative to the metaphase chromosomal aberration test as it examines cells at interphase and thus provides faster and less subjective results.22
Table 11.2 Some fish species with suitable karyotype for chromosomal aberration tests.

11.3.2 The Micronucleus Test
The micronucleus (MN) test was first introduced by Evans et al.23 in radiation-exposed Vicia faba root tips and subsequently developed and improved by several authors.24–26 Today, MN test has become a well-defined genotoxicity endpoint used in the assessment of structural and numerical chromosomal damages in different types of cells. The MN test is based on the detection of small membrane-bound DNA fragments (micronuclei) in interphase cells to measure structural and numerical chromosomal aberrations (Figure 11.3).
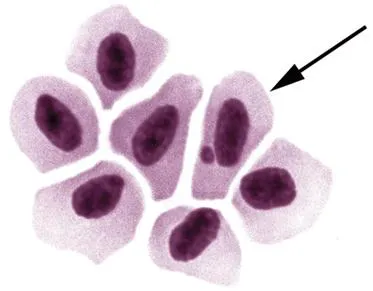
Figure 11.3 Photomicrograph of a micronucleated erythrocyte of O. niloticus (1000×; photo by T. Cavas).
A MN is composed of small chromatin fragments that arise as a result of chromosome breaks after clastogenic action or whole chromosomes that do not migrate during the anaphase as a result of aneugenic effects.27 These lagged chromosome fragments or chromosomes are enclosed by a nuclear membrane at the end of cell division and can be visualized by conventional staining in interphase cells (Figure 11.4). As a MN is originated during cell division, at least one cycle of cell division is necessary for the expression of MN in interphase cells.
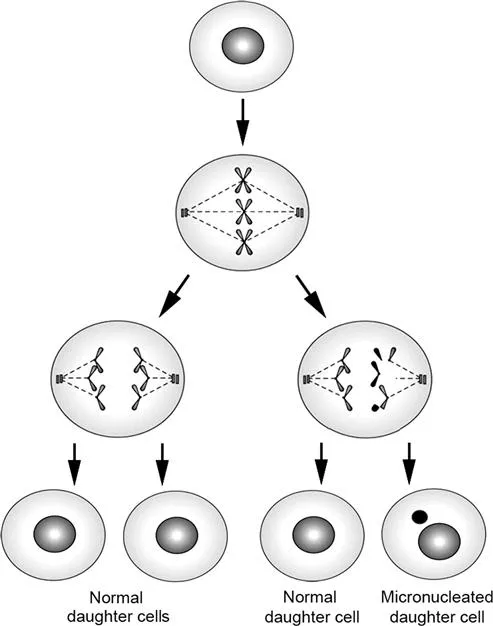
Figure 11.4 A schematic illustration of the mechanism of micronucleus formation in mononucleated cells following genotoxic exposure. Reprinted from Mutat. Res., 343, K. Al-Sabti and C. D. Metcalfe, Fish micronuclei for assessing genotoxicity in water, 121–135, Copyright (1995), with permission from Elsevier.
The criteria used for the identification of cells containing MN are as follows: (a) the diameter of the MN should be smaller than one-third of the main nucleus; (b) the MN should be clearly separated from the main nucleus, unless there is a clear identification of nuclear boundaries; and (c) the MN should not be refractive and should have a similar color and staining pattern as the main nucleus.
MN induction in fish was first reported by Hooftman and de Raat28 in peripheral erythrocytes of eastern mudminnow (Umbra pygmaea) treated with ethyl methanesulfonate. Furthermore, Al-Sabti29 demonstrated induction of micronuclei in three cyprinid fish following exposure to known mutagen substances. On the other hand, Manna and Sadhukhan30 presented a methodology for the micronucleus test in gill and kidney cells of tilapia (Oreochromis mossambicus) treated with X-rays and cadmium chloride. Furthermore, Williams and Metcalfe31 developed an in vivo hepatic micronucleus assay protocol with rainbow trout (Oncorhynchus mykiss). The possibility of the use of fin cells, which are in direct contact with water, was first demonstrated by Arkhipchuk and Garanko32 on common carp (Cyprinus carpio). Furthermore, Faßig;bender and Braunbeck5 successfully applied the micronucleus test on gonad cells of zebrafish (Danio rerio) exposed to methyl methanesulfonate.
Giemsa is the most commonly used staining technique in MN tests worldwide. However, in this staining method used in bright-field microscopy, some non-nucleic artifacts can also be stained and these artifacts could be mistakenly recorded as micronuclei. AO staining is better adapted to piscine erythrocyte MN assays than classical Giemsa staining. On the other hand, DNA-specific fluorescent staining reduces false-positive MN scoring due to artefacts. It was suggested that this specificity of OA provides more reliable and error-free scoring of MN. The acridine orange (AO) fluorescent staining method has been widely used in mammalian erythrocyte MN tests to discriminate immature polychromatic (PCE) and mature normochromatic (NCE) erythrocytes.33 AO is a nucleic acid-specific fluorochrome that stains DNA bright yellowish-green and RNA orange/reddish. Under a fluorescent microscope, micronuclei and main nuclei exhibit strong yellow-green fluoresc...
Table of contents
- Cover
- Title
- Copyright
- Preface
- Contents
- Section I: Aquatic Invertebrates as Experimental Models
- Section II: Aquatic Vertebrates as Experimental Models
- Subject Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Ecotoxicology and Genotoxicology by Marcelo L Larramendy in PDF and/or ePUB format, as well as other popular books in Physical Sciences & Ecology. We have over 1.5 million books available in our catalogue for you to explore.