Comprehensive coverage of the many different aspects of systems biology, resulting in an excellent overview of the experimental and computational approaches currently in use to study biological systems.
Each chapter represents a valuable introduction to one specific branch of systems biology, while also including the current state of the art and pointers to future directions. Following different methods for the integrative analysis of omics data, the book goes on to describe techniques that allow for the direct quantification of carbon fluxes in large metabolic networks, including the use of 13C labelled substrates and genome-scale metabolic models. The latter is explained on the basis of the model organism Escherichia coli as well as the human metabolism. Subsequently, the authors deal with the application of such techniques to human health and cell factory engineering, with a focus on recent progress in building genome-scale models and regulatory networks. They highlight the importance of such information for specific biological processes, including the ageing of cells, the immune system and organogenesis. The book concludes with a summary of recent advances in genome editing, which have allowed for precise genetic modifications, even with the dynamic control of gene expression.
This is part of the Advances Biotechnology series, covering all pertinent aspects of the field with each volume prepared by eminent scientists who are experts on the topic in question.

eBook - ePub
Systems Biology
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
About this book
Trusted by 375,005 students
Access to over 1 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Chapter 1
Integrative Analysis of Omics Data
Tobias Österlund, Marija Cvijovic and Erik Kristiansson
Summary
Data generation and analysis are essential parts of systems biology. Today, large amounts of omics data can be generated fast and cost-efficiently thanks to the development of modern high-throughput measurement techniques. Their interpretation is, however, challenging because of the high dimensionality and the often substantial levels of noise. Integrative analysis provides a framework for analysis of the omics data from a biological perspective, starting from the raw data, via preprocessing and statistical analysis, to the interpretation of the results. By integrating the data into structures created from biological information available in resources, databases, or genome-scale models, the focus moves from the individual transcripts or proteins to the entire pathways and other relevant biochemical functions present in the cell. The result provides a context-based interpretation of the omics data, which can be used to form a holistic and unbiased view of biological systems at a molecular level. The concept of integrative analysis can be used for many forms of omics data, including genome sequencing, transcriptomics, and proteomics, and can be applied to a wide range of fields within the life sciences.
1.1 Introduction
Systems biology is an interdisciplinary approach to biology and medicine that employs both experimentation and mathematical modeling to achieve a better understanding of biological systems by describing their shape, state, behavior, and evolutionary history. An important aim of systems biology is to deliver predictive and informative models that highlight the fundamental and presumably conserved relationships of biomolecular systems and thereby provide an improved insight into the many cellular processes [1]. Systems biology research methodology is a cyclical process fueled by quantitative experiments in combination with mathematical modeling (Figure 1.1) [2, 3]. In its most basic form, the cycle starts with the formulation of a set of hypotheses, which is followed by knowledge generation and model construction where an abstract description of the biological system (a model) is formulated and its parameters are estimated from data taken from the literature. The final step is defined by model predictions, where the constructed model is used to address the original hypotheses by providing a quantitative analysis of the system, which, in turn, generates new biological insight.
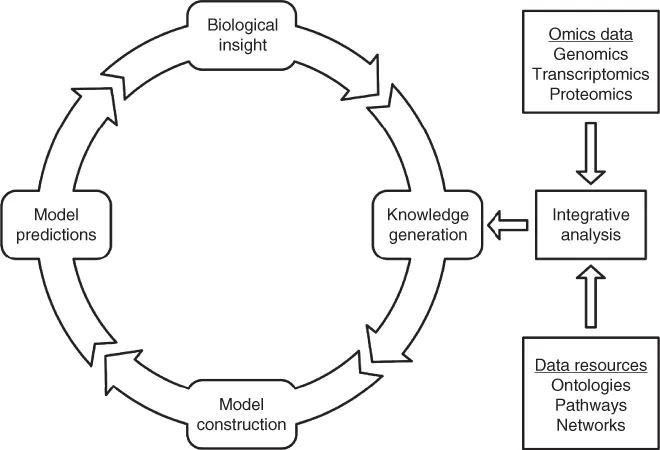
Figure 1.1 Systems biology research methodology. In the systems biology cycle, novel hypotheses are first formulated, which is followed by knowledge generation, model construction, and model predictions, which, in turn, leads to new biological insights. The development of high-throughput techniques have enabled rapid and cost-efficient generation of omics data from, for example, genome sequencing, transcriptomics, and proteomics. Integrative analysis provides a framework where omics data is systematically analyzed in a biological context, by data integration into known biological networks or other data resources, which enables improved interpretation and easier integration into quantitative models.
The development of high-throughput measurement techniques in the recent years has resulted in an unprecedented ability to rapidly and cost efficiently generate molecular data. Bioassays are today established for large-scale characterization of genes and their expression at the different layers defined by the central dogma: the genome, the transcriptome, and the proteome. The resulting data, which in this chapter will be referred to as omics data, is however complex because of its high dimensionality and is therefore hard to interpret and directly integrate into quantitative models. The concept of integrative analysis is a framework to systematically analyze the different components of omics data in relation to their corresponding biological functions and properties. The resulting biological interpretation can be used to form a holistic and unbiased view of biological systems at a molecular level. Thanks to the comprehensiveness of the omics data, all components (i.e., genes, transcripts, or proteins) can be measured simultaneously, which opens up opportunities for testing of existing hypotheses as well as generation of completely new hypotheses of the studied biological system.
The process of integrative analysis can be divided into two main steps: data processing and data integration (Figure 1.2). Integrative analysis starts from raw omics data and ends with the biological interpretation, and during this process the dimensionality of the data is reduced. The first step, the data processing, takes the high-dimensional omics data, and by applying computational and statistical tools, removes noise and errors while identifying genes and other components that contain information significant for the experiment. The next step, the data integration, uses the list of identified genes to pinpoint relevant functions and pathways by integrating the data on top of a “scaffold” built using established biological information collected from various resources and databases. The result, which is based on the combined analysis of the genes with similar functional properties, has a substantially reduced dimension, which considerably facilitates its interpretation.
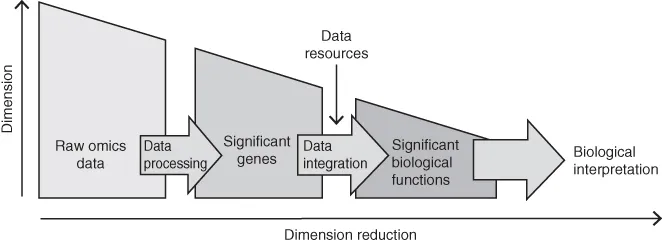
Figure 1.2 Description of the concept of integrative analysis as a tool for reduction of the dimension of omics data. Integrative analysis starts with raw omics data, which is typically affected by high levels of noise and errors. Computational and statistical approaches are first used to process the data to produce a ranked list of genes that are found to be of significant importance in the experiment. The gene list is used as input to the data integration, where known biological information is used as a basis for the interpretation of the data. During integrative analysis, the dimension of the data is significantly reduced, from potentially millions of data points to a limited number of significant biological functions and pathways, which considerably facilitates the interpretation.
Many studies in the life sciences aim to understand biological systems, often in relation to a perturbation caused by, for example, disease, genetic variability, changes in environmental parameters, or other factors introduced through laboratory experiments. A commonly used measurement technique is transcriptomics, where the transcriptional response is analyzed and the genes that are diff...
Table of contents
- Cover
- Related Titles
- Title Page
- Copyright
- Table of Contents
- List of Contributors
- About the Series Editors
- Chapter 1: Integrative Analysis of Omics Data
- Chapter 2: 13C Flux Analysis in Biotechnology and Medicine
- Chapter 3: Metabolic Modeling for Design of Cell Factories
- Chapter 4: Genome-Scale Metabolic Modeling and In silico Strain Design of Escherichia coli
- Chapter 5: Accelerating the Drug Development Pipeline with Genome-Scale Metabolic Network Reconstructions
- Chapter 6: Computational Modeling of Microbial Communities
- Chapter 7: Drug Targeting of the Human Microbiome
- Chapter 8: Toward Genome-Scale Models of Signal Transduction Networks
- Chapter 9: Systems Biology of Aging
- Chapter 10: Modeling the Dynamics of the Immune Response
- Chapter 11: Dynamics of Signal Transduction in Single Cells Quantified by Microscopy
- Chapter 12: Image-Based In silico Models of Organogenesis
- Chapter 13: Progress toward Quantitative Design Principles of Multicellular Systems
- Chapter 14: Precision Genome Editing for Systems Biology – A Temporal Perspective
- Index
- End User License Agreement
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.4M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Systems Biology by Jens Nielsen, Stefan Hohmann, Jens Nielsen,Stefan Hohmann, Sang Yup Lee,J. Nielsen,Gregory Stephanopoulos in PDF and/or ePUB format, as well as other popular books in Computer Science & Bioinformatics. We have over one million books available in our catalogue for you to explore.