
- 624 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
About this book
Pathogens respond dynamically to their environment. Understanding their behaviour is critical both because of evidence of increased resistance to established sanitation and preservation techniques, and because of the increased use of minimal processing technologies which are more vulnerable to the development of resistance. Understanding pathogen behaviour summarises the wealth of recent research and its implications for the food industry.After two introductory chapters on ways of analysing and modelling pathogens, Part one summarises current research on what determines pathogenicity, stress response, adaptation and resistance. Part two reviews the behaviour of particular pathogens, reviewing virulence, stress response and resistance mechanisms in such pathogens as Salmonella, E.coli and Campylobacter. The final part of the book assesses how pathogens react and adapt to particular stresses from heat treatment and the effects of low temperature to the use of disinfectants and sanitisers.With its distinguished editor and international team of contributors, Understanding pathogen behaviour is a standard reference for the food industry in ensuring food safety.
- Summarises the wealth of recent research in pathogen behaviour
- Assesses implications for microbiologists and QA staff in the food industry
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Part I
Understanding virulence, stress response and resistance mechanisms
1
Understanding the behaviour of pathogenic cells: proteome and metabolome analyses
S. Vaidyanathan and R. Goodacre, The University of Manchester, UK
1.1 Introduction
Post-genome science and technology are defined by the need to characterize function at the level of the genes, transcripts, proteins and metabolites to explain cellular processes. The understanding of biological systems is increasingly being driven by a paradigm shift in emphasis from the traditional divisive biochemical approaches that concentrate on local cellular processes, one at a time, to global approaches of analysing cellular compositions in parallel and in its entirety, with a view to obtain a more ‘holistic’ picture. Genomic sequencing initiatives have resulted in the sequencing of over 250 organisms, which include several pathogens (http://www.genomesonline.org). However, the functions of many (typically ~40 %) open reading frames (ORFs) within the sequenced genomes are still unknown. In this regard, analysis at the level of the functional units, i.e. transcripts (transcriptome), proteins (proteome) and metabolites (metabolome) is increasingly becoming relevant. In particular, post-transcriptional regulation of cellular activities necessitates analysis at the level of the proteome and the metabolome to better understand cellular processes. Strategies for the simultaneous high-throughput measurement of several analytes at the level of the proteome and metabolome are progressively being developed, and these form the subject-matter of this chapter.
In some cases, techniques that enable differences to be delineated between different biological systems or even between different states of a system may be useful in understanding cellular processes, even when a complete knowledge of the genetic make-up of the system is not available. In this regard, fingerprinting approaches are discussed that provide proteomic and metabolomic ‘snapshots’ and offer potential for rapid assessment of biological systems at the functional level. Although deductive reasoning dictates much of scientific inference, the practice of ‘holistic’ science and the inundation of data as a result necessitate a greater participation of inductive reasoning in the cycle of knowledge (Oldroyd, 1986) (Fig. 1.1). In this regard, computational sciences have much to offer, resulting in the birth of data-driven sciences such as ‘bioinformatics’, the application of which to proteome and metabolome analyses will also be discussed. Finally, the contribution of proteome and metabolome analyses to the understanding of pathogen behaviour is discussed, with relevant examples. The topical nature of the subject and its wide scope makes it difficult for a comprehensive coverage. Instead an attempt is made at capturing the general themes and trends to help keep the readers abreast of the developments.

1.2 Rationale behind analysing proteomes and metabolomes
The first complete genomic sequencing of a free-living organism was that of a human pathogen, Haemophilus influenzae, in 1995 (Fleischmann et al., 1995). Ever since, the genomic sequence of several pathogens have been revealed, including foodborne pathogens, with Campylobacter jejuni being the first food pathogen genome to be sequenced in 2000 (Parkhill et al., 2000). The availability of genetic information has enabled comparative genomic assessments that have contributed to the understanding of pathogen behaviour (Alm et al., 1999). However, genomic information alone is not sufficient for understanding biological processes. With the sequencing of the human genome, it is now known that we as humans have only three times as many genes as a nematode (Caenorhabditis elegans), with respect to the number of genes carried in our respective genetic make-up, and that the genetic make-up of diverse species are remarkably similar. Indeed genome sequencing projects alone have shown that we have 50 % genes in common with the fruit fly, 85 % with our canine friends and 99 % with chimpanzees; not forgetting that there is a significant portion shared with prokaryotes. This suggests that more than the genetic make-up, the contextual combination of gene products confers complexity and diversity to the functional genome. Consequently, in addition to cataloguing genomes and their function, it is necessary to generate an understanding of which gene products are expressed and how they come together to constitute a functional unit that responds to the different stimuli, be it of growth or environmentally induced. There is therefore a greater need to analyse at the level of the transcriptomes, proteomes and metabolomes (Fig. 1.2).
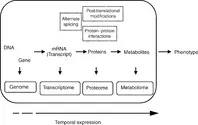
Transcriptomic analysis results in monitoring gene expression. Nucleic acid arrays produced by the robotic deposition of polymerase chain reaction (PCR) products, plasmids or oligonucleotides onto a glass slide or in situ synthesis of oligonucleotides using photolithography have been used in hybridization experiments to monitor gene expression. Array-based approaches, especially those that probe tens of thousands of genes, are useful in that they enable the development of a ‘holistic’ and unbiased view, rather than a targeted view of cellular response, without a priori knowledge of which genes or mechanisms are important. These and other tools, such as sequential analysis of gene expression (SAGE), can be used for monitoring morphological and physiological/phenotypical differences and can be indicative of cellular response to environmental stimuli and perturbations. However, mRNA is only an intermediate in translating the genetic information to cellular response and function, the business end of which is enacted by proteins and metabolites. Changes in the temporal expression and accumulation patt...
Table of contents
- Cover image
- Title page
- Table of Contents
- Related titles from Woodhead’s food science, technology and nutrition list
- Copyright
- Contributor contact details
- Introduction
- Part I: Understanding virulence, stress response and resistance mechanisms
- Part II: Virulence and stress response mechanisms of particular pathogens
- Part III: Pathogen resistance and adaptation to particular stresses
- Part IV: Appendix
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Understanding Pathogen Behaviour by M. Griffiths in PDF and/or ePUB format, as well as other popular books in Technology & Engineering & Food Science. We have over 1.5 million books available in our catalogue for you to explore.