
- 568 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Foodborne Infections and Intoxications
About this book
The accelerated globalization of the food supply, coupled with toughening government standards, is putting global food production, distribution, and retail industries under a high-intensity spotlight. High-publicity cases about foodborne illnesses over recent years have heightened public awareness of food safety issues, and momentum has been building to find new ways to detect and identify foodborne pathogens and eliminate food-related infections and intoxications. This extensively revised 4e covers how the incidence and impact of foodborne diseases is determined, foodborne intoxications with an introduction noting common features among these diseases and control measures that are applicable before and after the basic foodstuff is harvested.
- Provides a summary of the
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Subtopic
ToxikologieSection 1
Foodborne Disease: Epidemiology and Disease Burden
Chapter 1 Estimates of Disease Burden Associated with Contaminated Food in the United States and Globally
Chapter 2 The Foods Most Often Associated with Major Foodborne Pathogens
Chapter 3 Microbial Food Safety Risk Assessment
Chapter 4 Development of Risk-based Food Safety Systems for Foodborne Infections and Intoxications
Chapter 1
Estimates of Disease Burden Associated with Contaminated Food in the United States and Globally
Elaine Scallan1, Martyn Kirk2 and Patricia M. Griffin3, 1Colorado School of Public Health, Aurora, CO, USA, 2The Australian National University, Canberra, ACT, Australia, 3Enteric Diseases Epidemiology Branch, Division of Foodborne, Waterborne, and Environmental Diseases, Centers for Disease Control and Prevention, Atlanta, GA, USA
Introduction
Estimates of the overall burden of disease from foodborne agents are important for directing food safety policy and prioritizing interventions. However, estimating the burden of foodborne disease is challenging for several reasons. First, there are over 250 agents, including a variety of bacteria, viruses, parasites, and chemicals, that may contaminate food and cause foodborne illness. Second, transmission routes other than contaminated food may result in human infections for many of these agents. For example, Escherichia coli O157:H7 infections may be acquired by ingesting contaminated food or water or by direct contact with infected animals or persons. Third, a fraction of illnesses are confirmed by laboratory testing and reported to public health agencies and most surveillance systems do not attempt to determine the proportion of infections that are transmitted through food. Finally, unknown or unrecognized agents are likely to cause an important additional fraction of illnesses due to contaminated food. Indeed, many important foodborne pathogens, such as Campylobacter and E. coli O157, were only recognized in recent decades [1,2].
Surveillance for laboratory-confirmed infections provides essential information for assessing trends in diseases and detecting outbreaks. Information derived from surveillance may assist regulatory efforts to prioritize and evaluate interventions. However, because only a fraction of illnesses are diagnosed and reported, periodic assessments of the total number of illnesses, including those that are not laboratory-confirmed, are also needed to help set public health goals, allocate resources, and measure the economic impact. Several countries, including Australia, the Netherlands, the United Kingdom, and the United States, have conducted prospective population-based or cross-sectional studies to supplement surveillance and estimate the overall human health impact of foodborne disease [3]. In 2006, the World Health Organization (WHO) convened a meeting of foodborne disease experts that recommended the formation of the Foodborne Disease Epidemiology Reference Group (FERG) to advise WHO about how to estimate the global burden of foodborne disease [4]. The FERG began estimating the global burden of foodborne disease in 2007.
The purpose of this chapter is to describe the methods used by various countries to estimate the burden of foodborne disease. We begin by describing estimates of foodborne illness in the United States and then compare these methods and estimates with those in some other countries. We also discuss the WHO FERG initiative to estimate the global burden of foodborne disease.
Estimates of foodborne disease in the United States
In 2011, the US Centers for Disease Control and Prevention (CDC) published new estimates of the numbers of foodborne illnesses caused by contaminated foods consumed in the United States (hereafter, domestically acquired foodborne illnesses) [5,6]. Together, major known pathogens and unspecified agents transmitted by food were estimated to cause 47.8 million illnesses each year, resulting in 127,839 hospitalizations and 3037 deaths (Table 1.1).
Table 1.1
Estimated Annual Number of Illnesses, Hospitalizations, and Deaths Caused by Major Known Pathogens and Unspecified Agents Transmitted by Food (United States) [5,6].
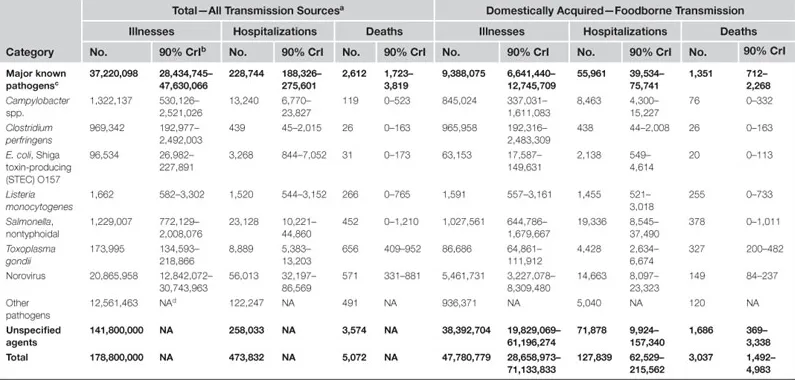
aIncludes all possible sources of illness, including infections acquired overseas.
b90% credible interval indicating range of uncertainty around estimate.
cShowing estimates for seven pathogens causing most foodborne illnesses, hospitalizations, or deaths.
dNA = not available.
Major known pathogens
Data from surveillance, surveys, and other sources were used to estimate the number of domestically acquired foodborne illnesses, hospitalizations, and deaths caused by 31 major known pathogens, including 21 bacterial, 5 viral, and 5 parasitic pathogens (see Table 1.1) [6]. These known pathogens were estimated to cause 9.4 million (90% credible interval [CrI]: 6.6–12.7 million) domestically acquired foodborne illnesses, 55,961 hospitalizations (90% CrI: 39,534–75,741), and 1351 deaths (90% CrI: 712–2268) each year. Norovirus was estimated to cause the most foodborne illness (58%), while nontyphoidal Salmonella spp. was the leading cause of hospitalization (35%) and death (28%). Seven pathogens—Campylobacter spp., Clostridium perfringens, E. coli O157, Listeria monocytogenes, nontyphoidal Salmonella spp., norovirus, and Toxoplasma gondii—were estimated to cause 90% of domestically acquired foodborne illnesses, hospitalizations, and deaths due to the major known pathogens.
Estimating illness using the “burden-of-illness pyramid”
Most known pathogens had laboratory-based surveillance data available; therefore, the total number of illnesses was estimated using the “burden-of-illness pyramid” approach (Figure 1.1). Several steps are necessary for an illness to be included in laboratory-based surveillance: the ill person must seek medical care, a specimen must be submitted for laboratory testing, the laboratory must test for and identify the causative agent, and the illness must be reported to the public health authorities. Estimating the frequency of cases of foodborne disease that are not reported to public health from laboratories provides insight into under-reporting in the surveillance system. Similarly, assessing differences in medical care–seeking behavior, specimen submission, laboratory testing, or laboratory test sensitivity characterizes under-diagnosis at each step of the surveillance system. Accounting for the proportion of cases missed in traditional surveillance due to under-diagnosis and under-reporting builds the “burden-of-illness pyramid”. This allows for an extrapolation from laboratory-confirmed illnesses (at the top of the “burden-of-illness pyramid”) to estimate the overall number of illnesses in the community (at the bottom of the “burden-of-illness pyramid”). To extrapolate, a multiplier, the inverse of a proportion, is calculated for each surveillance step. For example, if the laboratory test sensitivity of a particular pathogen was estimated to be 80%, the multiplier for this surveillance step would be 1.25 (i.e., for every case of infection diagnosed an estimated 1.25 cases would have been tested for that pathogen).
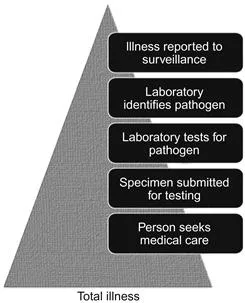
Figure 1.1 Surveillance steps that must occur for laboratory-confirmed cases to be reported to surveillance.
In the United States, data on laboratory-confirmed illnesses caused by 25 of the 31 known pathogens were available from one or more of five surveillance systems: the foodborne diseases active surveillance network (FoodNet), the national notifiable disease surveillance system (NNDSS), the cholera and other vibrio illness surveillance (COVIS) system, the national tuberculosis surveillance system (NTSS), and the foodborne disease outbreak surveillance system (FDOSS). Similar to other countries, laboratory-based surveillance systems in the United States rely largely upon passive reports of diseases from clinical laboratories to state and local health departments, which are, in turn sent to the CDC. To assure that all laboratory-confirmed cases occurring within the FoodNet surveillance area are reported, personnel actively contact all laboratories in the catchment area. Therefore, when data were available in more than one surveillance system, active surveillance data from FoodNet were used, except for Vibrio spp., for which COVIS was used because of geographical clustering of Vibrio infections outside the FoodNet sites. Data on outbreak-associated illnesses from FDOSS were used only for pathogens with no data available from the other systems due to not being specifically reported or only manifesting as outbreaks.
Because FoodNet conducts active surveillance, the pathogens under FoodNet surveillance were assumed to have no under-reporting. Because COVIS and NNDSS are passive surveillance systems, an under-reporting multiplier (1.1 for bacterial and 1.3 for parasitic pathogens), derived by comparing the incidence of all nationally notifiable illnesses ascertained through FoodNet with that reported to NNDSS, was applied to those pathogen counts. For the five bacterial pathogens for which only outbreak data were available, an outbreak under-reporting multiplier was created by determining the proportion of illnesses in FoodNet caused by Campylobacter, Cryptosporidium, Cyclospora, Listeria, Salmonella, Shigella, Shiga toxin-producing E. coli (STEC), Vibrio, and Yersinia that were also reported as outbreaks associated to FDOSS. It was assumed that all Mycobacterium bovis illnesses were reported to NTSS.
To adjust for medical care seeking and specimen submission, the proportion of persons reporting an acute diarrheal illness (defined as ≥3 loose stools in a 24-hour period and lasting longer than one day or resulting in restricted daily activities) in the past month who sought medical care and submitted a stool sample for that illness were estimated using data from FoodNet surveys of the general population (FoodNet Population Surveys). Because persons with more severe illness are more likely to seek care [7], the rate of medical care seeking and stool sample submission was estimated separately for persons with bloody and non-bloody diarrhea; these proportions were used as surrogates for severe and mild presentations of most illnesses. These multipliers were derived by examining data on the proportion of patients with diarrhea seeking care and submitting specimens with different symptom profiles from population-based surveys. Multipliers for medical care seeking and stool sample submission (for those with mild and severe illness) were then appli...
Table of contents
- Cover image
- Title page
- Table of Contents
- Copyright
- List of Contributors
- Preface
- Preface to the Third Edition
- Dedication
- Section 1: Foodborne Disease: Epidemiology and Disease Burden
- Section 2: Foodborne Infections: Bacterial
- Section 3: Foodborne Infections: Viral
- Section 4: Foodborne Infections: Parasites and Others
- Section 5: Intoxications
- Section 6: Policy and Prevention of Foodborne Diseases
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Foodborne Infections and Intoxications by J. Glenn Morris Jr. in PDF and/or ePUB format, as well as other popular books in Technik & Maschinenbau & Toxikologie. We have over 1.5 million books available in our catalogue for you to explore.